Dr Redmond Smyth, Former Group Leader
Genome Architecture and Evolution of RNA viruses
From 2018 to 2024, a research group at the Helmholtz Institute Würzburg (HIRI), led by Redmond Smyth, focused on the genome architecture and evolution of RNA viruses. While maintaining close ties to research at the HIRI, Smyth now leads a research group in Strasbourg, France, where he was appointed by the Centre national de la recherche scientifique (CNRS) in 2024.
Our research and approach
RNA viruses are an important class of emerging infectious diseases. Currently, the majority of antivirals and pharmaceuticals function at protein level. However, RNA structures within viral genomes interact with cellular/viral proteins and nucleic acids to provide a novel surface for pharmaceutical intervention. A detailed understanding of these processes is necessary for the development of novel RNA-based therapeutics.
Redmond Smyth's group studied RNA-based mechanisms of RNA virus replication and evolution. Their work employed a combination of molecular virology, RNA biochemistry and mathematical modeling. They sought to identify essential RNA structures as new antiviral targets, and to understand how RNA structure regulates viral evolution for better vaccine and pandemic preparedness.
Their investigations focused on single molecule and single virion analyses using high-throughput, genome-wide chemical probing technologies and mutation based functional screens. They also developed new technologies including RNA-RNA-seq, single-molecule RNA structural probing and sequencing coupled microscopy. The group's overarching goal was to exploit knowledge of RNA virus genome architecture to advance RNA-based therapies.
Team members

Dr Redmond Smyth, Former Group Leader
Group Leader
Research projects
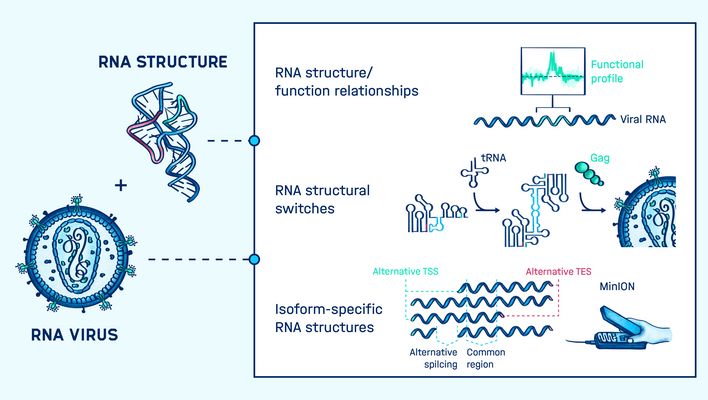
Graphical Abstract
Alumni
Shazeb Ahmad, PhD Student • Uddhav Ambi, PhD Student • Patrick Bohn, PhD Student • Charlene Börtlein, Technical Assistant • Anne-Sophie Gribling, Postdoc • Jianhui Li, PhD Student • Marco Antonio Olguín Nava, PhD Student • Liqing Ye, PhD Student
Publications
2025
The translational landscape of HIV-1 infected cells reveals key gene regulatory principles
Kibe A, Buck S, Gribling-Burrer AS, Gilmer O, Bohn P, Koch T, Mireisz CN, Schlosser A, Erhard F, Smyth RP, Caliskan N (2025)
Nature Structural & Molecular Biology 32 (5): 841-852
High-Fidelity and Cost-Effective Engineering of SARS-CoV-2
Olguin-Nava M, Hennig T, Börtlein C, Bohn P, Ambi UB, Gabel A, Günter LM, Gribling-Burrer AS, Schmidt N, Caliskan N, …, Munschauer M, Smyth RP (2025)
Viruses 17 (12): 1604
Cell type-specific purifying selection of synonymous mitochondrial DNA variation
Lareau CA, Maschmeyer P, Yin Y, Gutierrez JC, Dhindsa RS, Gribling-Burrer AS, Zielinski S, Hsieh YH, Nitsch L, Dimitrova V, …, Ludwig LS, Satpathy AT (2025)
PNAS 122 (30): e2505704122
Visualizing the transcription and replication of influenza A viral RNAs in cells by multiple direct RNA padlock probing and in situ sequencing (mudRapp-seq)
Ahmad S, Li J, Schaust J, Gribling-Burrer AS, Geiger N, Fischer SC, Ambi UB, Backes S, Ankenbrand MJ, Smyth RP (2025)
Nucleic Acids Research 53 (11): gkaf461
Demultiplexing and barcode-specific adaptive sampling for nanopore direct RNA sequencing
van der Toorn W, Bohn P, Liu-Wei W, Olguin-Nava M, Gribling-Burrer AS, Smyth RP, von Kleist M (2025)
Nature Communications 16 (1): 3742
Giant RNA genomes: Roles of host, translation elongation, genome architecture, and proteome in nidoviruses
Neuman BW, Smart A, Gilmer O, Smyth RP, Vaas J, Böker N, Samborskiy DV, Bartenschlager R, Seitz S, Gorbalenya AE, Caliskan N, Lauber C (2025)
PNAS 122 (7): e2413675122
Selectively expressed RNA molecules as a versatile tool for functionalized cell targeting
Rastfeld F, Hoffmann M, Krüger S, Bohn P, Gribling-Burrer AS, Wagner L, Hersch N, Stegmayr C, Lövenich L, Gerlach S, …, Rueger MA, Hoffmann B (2025)
Nature Communications 16 (1): 420
2024
Isoform-specific RNA structure determination using Nano-DMS-MaP
Gribling-Burrer AS, Bohn P, Smyth RP (2024)
Nature Protocols 19 (6): 1835-1865
The conserved noncoding RNA ModT coordinates growth and virulence in Clostridioides difficile
Lence T, Sulzer J, Andress K, Gribling-Burrer AS, Lamm-Schmidt V, Barquist L, Smyth RP, Faber F (2024)
PLOS Biology 22 (12): e3002948
NEAT1 promotes genome stability via m6A methylation-dependent regulation of CHD4
Mamontova V, Trifault B, Gribling-Burrer AS, Bohn P, Boten L, Preckwinkel P, Gallant P, Solvie D, Ade CP, Papadopoulos D, …, Smyth RP, Burger K (2024)
Genes & Development 38 (17-20): 915-930
Recruitment of multi-segment genomic RNAs by Bluetongue virus requires a preformed RNA network
Sung PY, Phelan JE, Luo D, Kulasegaran-Shylini R, Bohn P, Smyth RP, Roy P (2024)P., Roy P (2024)
Nucleic Acids Research 52 (14): 8500-8514
Sequencing accuracy and systematic errors of nanopore direct RNA sequencing
Liu-Wei W, van der Toorn W, Bohn P, Hölzer M, Smyth RP, von Kleist M (2024)
BMC Genomics 25 (1): 528
2023
Cis-mediated interactions of the SARS-CoV-2 frameshift RNA alter its conformations and affect function
Pekarek L, Zimmer MM, Gribling-Burrer AS, Buck S, Smyth RP, Caliskan N (2023)
Nucleic Acids Research 51 (2): 728–743
SND1 binds SARS-CoV-2 negative-sense RNA and promotes viral RNA synthesis through NSP9
Schmidt N, Ganskih S, Wei Y, Gabel A, Zielinski S, Keshishian H, Lareau CA, Zimmermann L, Makroczyova J, Pearce C, …, Erhard F, Munschauer M (2023)
Cell 186 (22): 4834-4850.e23
Advanced fluorescence microscopy in respiratory virus cell biology
Xie E, Ahmad S, Smyth RP, Sieben C (2023)
Advances in Virus Research 116: 123-172
Sequential disruption of SPLASH-identified vRNA-vRNA interactions challenges their role in influenza A virus genome packaging
Jakob C, Lovate GL, Desirò D, Gießler L, Smyth RP, Marquet R, Lamkiewicz K, Marz M, Schwemmle M, Bolte H (2023)
Nucleic Acids Research 51 (12): 6479-6494
Nano-DMS-MaP allows isoform-specific RNA structure determination
Bohn P, Gribling-Burrer AS, Ambi UB, Smyth RP (2023)
Nature Methods 20 (6): 849-859
2022
Short- and long-range interactions in the HIV-1 5' UTR regulate genome dimerization and packaging
Ye L, Gribling-Burrer AS, Bohn P, Kibe A, Börtlein C, Ambi UB, Ahmad S, Olguin-Nava M, Smith M, Caliskan N, von Kleist M, Smyth RP (2022)
Nature Structural & Molecular Biology 29 (4): 306-319
2021
The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting
Zimmer MM, Kibe A, Rand U, Pekarek L, Ye L, Buck S, Smyth RP, Cicin-Sain L, Caliskan N (2021)
Nature Communications 12 (1): 7193
RNA Structures and Their Role in Selective Genome Packaging
Ye L, Ambi UB, Olguin-Nava M, Gribling-Burrer AS, Ahmad S, Bohn P, Weber MM, Smyth RP (2021)
Viruses 13 (9): 1788
2019
The evolution of RNA structural probing methods: From gels to next-generation sequencing
Mailler E, Paillart J, Marquet R, Smyth RP, Vivet-Boudou V (2019)
Wiley Interdisciplinary Reviews: RNA 10 (2): e1518
2018
In cell mutational interference mapping experiment (in cell MIME) identifies the 5' polyadenylation signal as a dual regulator of HIV-1 genomic RNA production and packaging
Smyth RP, Smith MR, Jousset A, Despons L, Laumond G, Decoville T, Cattenoz P, Moog C, Jossinet F, Mougel M, …, Kleist M, Marquet R (2018)
Nucleic Acids Research 46 (9): e57
RNA Structure - A Neglected Puppet Master for the Evolution of Virus and Host Immunity
Smyth RP, Negroni M, Lever AM, Mak J, Kenyon JC (2018)
Frontiers in Immunology 9: 2097
Structural and Functional Motifs in Influenza Virus RNAs
Ferhadian D, Contrant M, Printz-Schweigert A, Smyth RP, Paillart J, Marquet R (2018)
Frontiers in Microbiology 9: 559
2017
HIV-1 Pr55Gag binds genomic and spliced RNAs with different affinity and stoichiometry
Bernacchi S, Abd El-Wahab EW, Dubois N, Hijnen M, Smyth RP, Mak J, Marquet R, Paillart J (2017)
RNA Biology 14 (1): 90-103
2016
HIV-1 Mutation and Recombination Rates Are Different in Macrophages and T-cells
Cromer D, Schlub TE, Smyth RP, Grimm AJ, Chopra A, Mallal S, Davenport MP, Mak J (2016)
Viruses 8 (4): 118
The Life-Cycle of the HIV-1 Gag-RNA Complex
Mailler E, Bernacchi S, Marquet R, Paillart J, Vivet-Boudou V, Smyth RP (2016)
Viruses 8 (9): E248
MIMEAnTo: profiling functional RNA in mutational interference mapping experiments
Smith MR, Smyth RP, Marquet R, Kleist M (2016)
Bioinformatics 32 (21): 3369-3370
2015
Mutational interference mapping experiment (MIME) for studying RNA structure and function
Smyth RP, Despons L, Huili G, Bernacchi S, Hijnen M, Mak J, Jossinet F, Weixi L, Paillart J, Kleist M, Marquet R (2015)
Nature Methods 12 (9): 866-72
Evaluation of anti-HIV-1 mutagenic nucleoside analogues
Vivet-Boudou V, Isel C, El Safadi Y, Smyth RP, Laumond G, Moog C, Paillart J, Marquet R (2015)
The Journal of Biological Chemistry 290 (1): 371-83
Properties of HIV-1 associated cholesterol in addition to raft formation are important for virus infection
Hawkes D, Jones KL, Smyth RP, Pereira CF, Bittman R, Jaworowski A, Mak J (2015)
Virus Research 210: 18-21
2014
Specific recognition of the HIV-1 genomic RNA by the Gag precursor
Abd El-Wahab EW, Smyth RP, Mailler E, Bernacchi S, Vivet-Boudou V, Hijnen M, Jossinet F, Mak J, Paillart J, Marquet R (2014)
Nature Communications 5: 4304
Identifying recombination hot spots in the HIV-1 genome
Smyth RP, Schlub TE, Grimm AJ, Waugh C, Ellenberg P, Chopra A, Mallal S, Cromer D, Mak J, Davenport MP (2014)
Journal of Virology 88 (5): 2891-902
Fifteen to twenty percent of HIV substitution mutations are associated with recombination
Schlub TE, Grimm AJ, Smyth RP, Cromer D, Chopra A, Mallal S, Venturi V, Waugh C, Mak J, Davenport MP (2014)
Journal of Virology 88 (7): 3837-49
2013
Improved quantification of HIV-1-infected CD4+ T cells using an optimised method of intracellular HIV-1 gag p24 antigen detection
Yang H, Yorke E, Hancock G, Clutton G, Sande N, Angus B, Smyth RP, Mak J, Dorrell L (2013)
Journal of Immunological Methods 391 (1-2): 174-8
Intracellular Dynamics of HIV Infection
Petravic J, Ellenberg P, Chan M, Paukovics G, Smyth RP, Mak J, Davenport MP (2013)
Journal of Virology 88 (2): 1113-24
A functional sequence-specific interaction between influenza A virus genomic RNA segments
Gavazzi C, Yver M, Isel C, Smyth RP, Rosa-Calatrava M, Lina B, Moulès V, Marquet R (2013)
PNAS 110 (41): 16604-9
2012
The Origin of Genetic Diversity in HIV-1
Smyth RP, Davenport MP, Mak J (2012)
Virus Research 169 (2): 415-29
2011
8-Modified-2'-deoxyadenosine analogues induce delayed polymerization arrest during HIV-1 reverse transcription
Vivet-Boudou V, Isel C, Sleiman M, Smyth RP, Ben Gaied N, Barhoum P, Laumond G, Bec G, Götte M, Mak J, …, Burger A, Marquet R (2011)
PLOS One 6 (11): e27456
Early events of HIV-1 infection: can signaling be the next therapeutic target?
Jones KL, Smyth RP, Pereira CF, Cameron PU, Lewin SR, Jaworowski A, Mak J (2011)
Journal of neuroimmune pharmacology : the official journal of the Society on NeuroImmune Pharmacology 6 (2): 269-83
Labeling of multiple HIV-1 proteins with the biarsenical-tetracysteine system
Pereira CF, Ellenberg PC, Jones KL, Fernandez TL, Smyth RP, Hawkes DJ, Hijnen M, Vivet-Boudou V, Marquet R, Johnson I, Mak J (2011)
PLOS One 6 (2): e17016
2010
Accurately measuring recombination between closely related HIV-1 genomes
Schlub TE, Smyth RP, Grimm AJ, Mak J, Davenport MP (2010)
PLOS Computational Biology 6 (4): e1000766
Reducing chimera formation during PCR amplification to ensure accurate genotyping
Smyth RP, Schlub TE, Grimm A, Venturi V, Chopra A, Mallal S, Davenport MP, Mak J (2010)
Gene 469 (1-2): 45-51
2009
The A-rich RNA sequences of HIV-1 pol are important for the synthesis of viral cDNA
Keating CP, Hill MK, Hawkes DJ, Smyth RP, Isel C, Le S, Palmenberg AC, Marshall JA, Marquet R, Nabel GJ, Mak J (2009)
Nucleic Acids Research 37 (3): 945-56