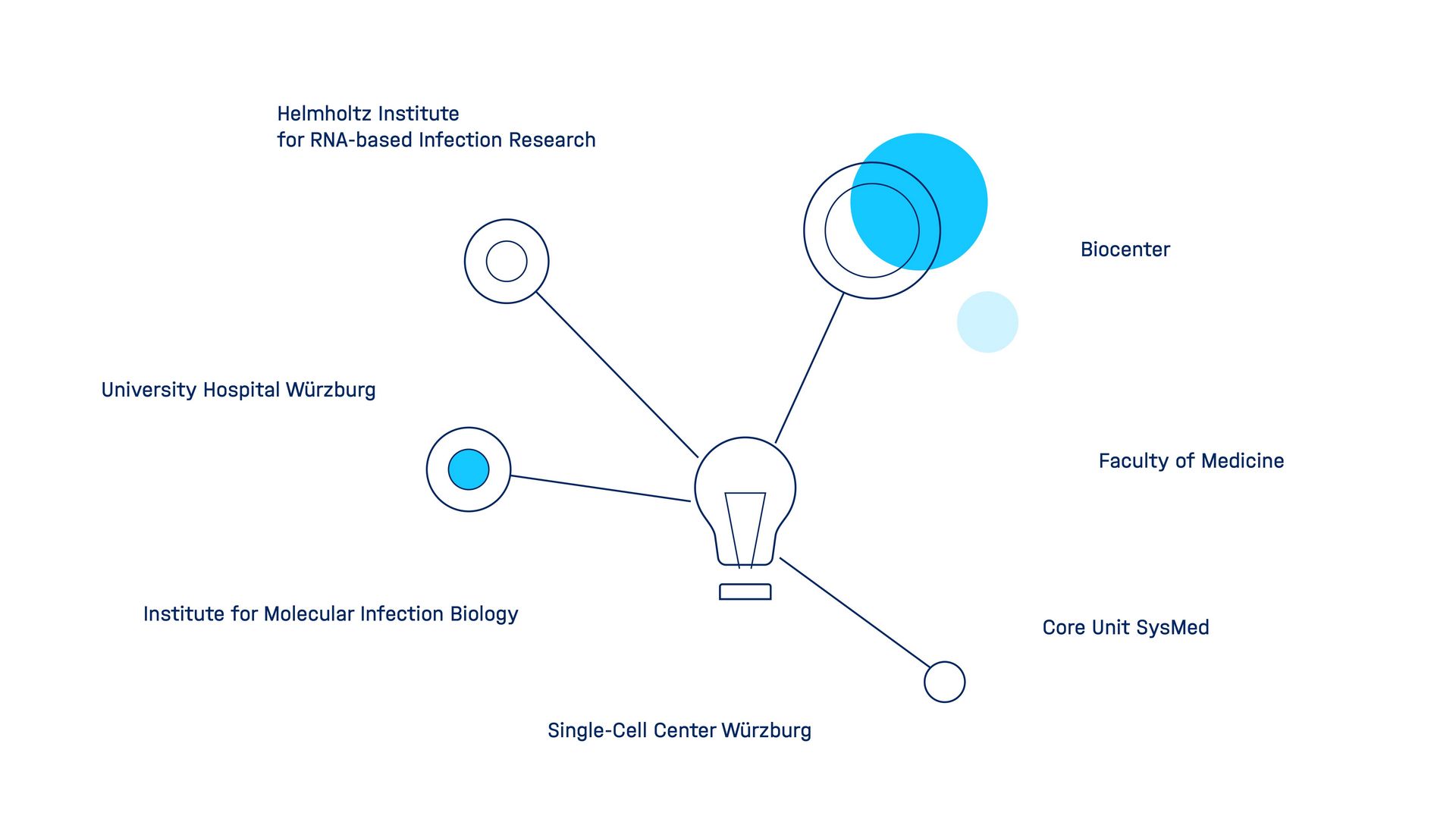
Würzburg RNA Ecosystem
The Würzburg RNA Ecosystem is characterized by interdisciplinary research and technology centers such as the Biocenter and the Single-Cell Center. The Helmholtz Institute for RNA-based Infection Research (HIRI) closely cooperates with various partners such as the Max Planck and Fraunhofer Societies to foster local interdisciplinary research.
Würzburg RNA Ecosystem: Institutions
Helmholtz Institute for RNA-based Infection Research
The Helmholtz Institute for RNA-based Infection Research (HIRI) was established in May 2017 as a site of the Helmholtz Centre for Infection Research (HZI) in Braunschweig in cooperation with the Julius-Maximilians-Universität of Würzburg (JMU). Located on the Würzburg medical campus, the HIRI is the first research institution of its kind worldwide to fully focus on the role of RNA in infection processes.
Rising antimicrobial resistance, chronic infections, and (re-)emerging pathogens are among the major challenges facing humanity. As a federal institute, the HIRI pioneers an integrative approach to exploit the vast potential of RNA as a diagnostic, drug, and therapeutic target for new strategies to combat infectious diseases.
Led by Director Jörg Vogel, the HIRI focuses on three central areas: Basic research on bacterial pathogens, on viruses and on the host response by the immune system. These three fields are complemented by applied research on diagnostics and RNA delivery for therapeutic purposes. The integration of emerging topics in RNA research, such as RNA modifications, will ensure a place for the HIRI at the forefront of infectious disease research. Core technologies, with a major focus on sequencing approaches, are harnessed for high-throughput exploration of RNA functions and will provide an opportunity to construct a systems-level view of the role of RNA during infection.
Integrated scientific concept
With its integrated scientific concept and cutting-edge infrastructure, the HIRI provides a vibrant research environment for both established and young scientists.
The natural synergy between this new institute and the infection research and translational competences at the University of Würzburg as well as at the Helmholtz Centre for Infection Research (HZI) Braunschweig creates unique opportunities to effectively convert knowledge into applications.
This setting also stimulates collaborations with industry in the development of novel therapeutic and diagnostic strategies for the treatment of infectious diseases.
Institute for Molecular Infection Biology
The Institute for Molecular Infection Biology (IMIB) at the Julius-Maximilians-Universität of Würzburg (JMU) is an interdisciplinary research institution of the Medical Faculty with strong links to the Faculty of Biology. Jörg Vogel, who is also director of the HIRI, has been director of the institute since 2009.
The internationally recruited scientists at IMIB investigate the molecular mechanisms of infections caused by bacteria, parasites and fungi. They aim to elucidate fundamental aspects of gene regulation of pathogens and to identify common strategies employed by the pathogens when interacting with host cells and the immune system.
Another focus of the institute lies in the analysis of small RNAs and other non-coding RNA molecules in infection. This is complemented by a research technology infrastructure that aims at developing new applications for sequencing-based approaches like single-cell RNA-seq, multi-organism-RNA-seq and
tissue RNA-seq.
Single-Cell Center Würzburg
In 2021, the HIRI co-founded the Single-Cell Center Würzburg to further advance one of its core technologies. The Center is a joint venture with the Faculty of Medicine of JMU, the University Hospital Würzburg (UKW), the Fraunhofer Translational Center for Regenerative Therapies (TLZ-RT), and the Max Planck Research Group at the Würzburg Institute of Systems Immunology (WüSI).
The center’s objective is to analyze and understand diseases at the level of individual cells. In the future, this will enable the earliest possible and most reliable prediction of a disease and how it can be treated in the best possible way. The center combines local scientific, clinical and methodological expertise relevant to innovative single-cell research and provides access to the full range of scRNA-seq technologies for both eukaryotes and microorganisms.
Starting from infection research, the Würzburg Single-Cell Center will also investigate cancer, neurodegenerative disorders or cardiovascular diseases at the single cell level. The broad method spectrum of RNA sequencing technologies is to be rapidly enhanced in this context.
Detect a disease before it emerges? This is how single-cell research may help:
Core Unit SysMed
The Core Unit Systems Medicine (SysMed) is a joint institution of the JMU Medical Faculty and the University Hospital. It works closely with the HIRI and other research institutes. It provides scientists with the technology and bioinformatics support for RNA and DNA sequencing for biomedical research. A particular focus of the unit is the application of single-cell RNA sequencing. The establishment of additional subunits for the support of large-scale data analyses and modelling are in progress.
Biocenter
The Biocenter is an amalgamation of departments and chairs from the Faculties of Biology, Medicine and Chemistry, collaborating in research and teaching. It unites scientists who share the conviction that the great questions of biology and medicine can only be answered if they are tackled on each of the organizational levels of biology − from molecules, cells, organs and the whole organism, to interactions between organisms.
Consequently, researchers at the Biocenter are developing the latest methods in molecular biology, cell biology and organismic biology and applying them closely to their respective scientific research.
Research is being done using almost all important model organisms of the animal and plant kingdoms, including Arabidopsis, Drosophila, honey bees, the mouse, xenopus, yeast and zebrafish. Technologies at the interface of several disciplines − such as advanced microscopy, High Throughput Screening, DNA sequencing and metabolome analysis − are organized as central facilities.
Medical Faculty (JMU)
The Medical Faculty of the Julius-Maximilians-Universität of Würzburg (JMU) has a strong connection to the University Hospital, founded on scientific research carried out in close cooperation with clinical institutes.
By transcending traditional clinical research areas and exchanging them for three novel complexity-oriented research foci the faculty aims to represent itself in its entirety more effectively, to overcome borders between traditional disciplines, and to foster interdisciplinarity. The three newly defined research foci are: (I) cellular heterogeneity, (II) complexity within organ tissues and (III) system and network diseases. Focusing on these research foci does not dispense from the necessity of excellent discipline-specific research; they rather provide a cross-disciplinary framework for joint projects that may eventuate in collaborative research initiatives.
Research within the Medical Faculty is structurally supported by platform technologies such as microscopy, mass spectrometry, transgene technologies, and systems medicine.
University Hospital Würzburg
The University Hospital Würzburg (UKW) is devoted to medical care, research and teaching, which facilitates progressive counselling and treatment of patients according to the highest current standards in medical care: First-class medicine and research that benefits all patients.
The hospital assumes a central role in providing high quality medical care to the people of Würzburg and the surrounding region. It consists of 19 clinics and three autonomous policlinics as well as four clinical institutes. Each year, UKW takes care of about 72,000 stationary patients, plus more than 264,000 outpatient treatments.
In addition, the hospital is involved in several clinical research centers, where interdisciplinary cooperation takes the center stage: Amongst others these are the Comprehensive Cancer Center Mainfranken, the Interdisciplinary Center for Palliative Medicine and the Comprehensive Heart Failure Center.
UKW is an important resource for many scientists providing them with tissue samples for basic research again highlighting the local scientific network and support structures.
RNA Faculty Members
Jörg Vogel

Jörg Vogel © Nik Schölzel
RNA Biology of Bacterial Infections
The research group led by Jörg Vogel investigates the RNA world of microbes during infection and in the context of their natural habitat. Using high-resolution methods, they seek to understand when, how and why bacteria use RNA-based mechanisms to regulate their genes, and exploit this knowledge to develop RNA-based therapeutic approaches.
Chase Beisel

Chase Beisel © Nik Schölzel
RNA Synthetic Biology
The research group led by Chase Beisel works at the interface of CRISPR biology and technologies. They investigate the functional diversity of CRISPR-Cas immune systems to develop new technologies that can be employed to better understand, diagnose and treat human infections.
Emmanuel Saliba

Emmanuel Saliba © Nik Schölzel
Single-Cell Analysis
The research group led by Emmanuel Saliba explores host-pathogen interactions in high resolution at the single-cell level. They develop and integrate single-cell genomics, imaging and computational approaches to decipher the microenvironments of individual pathogens and shed light on the heterogeneity of host responses and disease outcomes.
Jens Hör

Jens Hör © Luisa Härtig
Molecular Principles of RNA Phages
The research group led by Jens Hör investigates the dynamics of infection, host takeover, and anti-phage defense during the interaction between RNA phages and their hosts. They seek to understand the molecular principles underlying these processes to develop new and improved antibacterial strategies such as phage therapy.
Franziska Faber

Franziska Faber © Nik Schölzel
RNA Biology of Gram-positive Bacteria
The research group led by Franziska Faber investigates the interplay between the enteric pathogen Clostridioides difficile and the intestinal microbiota. They seek to better understand the RNA-based mechanisms controlling virulence during these interactions. The goal is to leverage this knowledge for the development of novel RNA-based antimicrobials.
Camilla Ciolli Mattioli

Camilla Ciolli Mattioli © privat
Systems Microbiology of Intracellular Pathogens
The research group led by Camilla Ciolli Mattioli decodes how intracellular bacteria, like Salmonella, survive, persist, and evade destruction within host cells. By revealing the molecular mechanisms that determine infection outcomes, the team aims to develop new strategies to combat antibiotic-resistant infections.
Utz Fischer

Utz Fischer © privat
RNA Metabolism and Macromolecular Machines
The Fischer group studies the functional dynamics of macromolecular machines acting on RNA using a combination of biochemical, structural and systems biology approaches. By combining basic RNA research with biomedical approaches, the scientists also investigate how RNA-based pathways and networks are changed in different disease settings and in viral infections.
Dominic Grün

Dominic Grün © Marcus Rockoff
Quantitative Single-Cell Biology of the Immune System
Cell fate decision is essential for the emergence of a complex multicellular organism starting from a single pluripotent zygote, and perpetually occurs during adult life to maintain organ tissues. The robustness of cell fate decision is remarkable given the stochastic interaction of hundreds of thousands of molecules in each cell. It has remained a central open question how signals from the microenvironment are integrated with stochastic processes in a cell to control cell fate decision in normal tissues and upon perturbations due to disease or tissue damage. The Grün lab investigates these questions in the context of immune cell differentiation by combining single-cell RNA-seq methods with computational approaches involving machine learning and mathematical modeling.
Claudia Höbartner

Claudia Höbartner © privat
Organic and Biomolecular Chemistry
- Chemical biology of natural RNA/DNA modifications
- Nucleic acid chemistry
- DNA catalysis
- Ribozymes and aptamers
- Biomolecular label chemistry
- Structure and mechanism of functional nucleic acids and RNA protein complexes
Julian König

Julian König © privat
RNA Modification and Regulation
The research group led by Julian König works on deciphering how RNA processing is controlled by RNA binding proteins and RNA modifications. By integrating transcriptomic, biochemical, and functional genomics approaches, the group seeks to decode the RNA regulatory code. Their ultimate goal is to translate insights from fundamental research into an understanding of human physiology and disease.
Katharina Markmann

Katharina Markmann © Daniel Peter
Small RNAs in systemic control of plant symbiosis and development
The research group led by Katharina Markmann seeks to understand the molecular mechanisms underlying systemic control of plant root endosymbioses with bacteria and fungi, and the role of regulatory small RNAs therein. They further analyze the role of systemically mobile small RNAs in controlling plant developmental responses to biotic and abiotic stimuli.
Lorenz Meinel

Lorenz Meinel © Ludwig Höllein
Drug Formulation and Delivery
Biological interfaces:
- Bioresponsive systems (diagnostics and therapy)
- Pharmaceutical functionalization of biomaterials
- Formulation of proteins, chemical drugs and therapeutic gases
Chemical-physical interfaces:
- Transformation of drugs into ionic liquids
Thomas Rudel

Thomas Rudel © Hilde Merkert
Bacteria-Host Interaction
The Rudel lab studies molecular mechanisms how the major human pathogenic bacteria Chlamydia trachomatis, Neisseria gonorrhoeae and Staphylococcus aureus cause diseases. The group’s research focuses on the interaction of these pathogens with different host cells and how this disturbs the normal function of the cell and the innate immune and cell autonomous defense. One particular aspect in this context is the role of bacterial (sRNAs) and human regulatory RNAs (microRNAs) during the infection process. The researchers aim to understand invasive (S. aureus sepsis; disseminating gonococcal infection) and chronic diseases (Chlamydia ovarian cancer connection) by employing new infection models on the basis of engineered human three dimensional tissues.
Cynthia Sharma

Cynthia Sharma © Petra Thomas
Deep Sequencing Approaches to Pathogenesis
The Sharma lab focuses on how bacterial pathogens adapt to changing environments and control their virulence. They aim to unravel mechanisms and functions of gene regulation via small RNAs (sRNAs) and associated RNA binding proteins (RBPs) as well as small proteins. Their main model organisms are the gastric pathogen Helicobacter pylori and the related foodborne pathogen Campylobacter jejuni. They have been developing and employing diverse deep-sequencing approaches (e.g. RNA-seq, Ribo-seq, RIP-seq) for global transcriptome and translatome analysis and to study RNA-protein complexes.
Using genomics, genetics, and biochemical approaches, they aim to identify and characterize RNA and protein factors, such as RBPs and ribonucleases, involved in post-transcriptional regulation in bacterial pathogens. Using three-dimensional (3D) infection models based on tissue engineering, they are investigating the roles of sRNAs, small proteins, and RBPs in virulence of H. pylori and C. jejuni. Moreover, they are exploring mechanisms and functions of bacterial CRISPR-Cas immune systems, leading to new biotechnological applications.
Alexander Westermann

Alexander Westermann © Nik Schölzel
Host-Pathogen-Microbiota Interactions
The research group led by Alexander Westermann investigates molecular RNA-based mechanisms underlying the interplay between the host, invading pathogens and the host microbiota. They seek to identify potential molecules and metabolic pathways to target in diagnostics and therapeutics.
Kathi Zarnack

Kathi Zarnack © Robert Emmerich / Uni Würzburg
Bioinformatics of RNA Regulation and Gene Expression
The Zarnack group focuses on understanding the molecular mechanisms of RNA regulation in human cells and how their dysregulation contributes to disease. Using transcriptomic data and advanced bioinformatics, we explore the intricate interplay of RNA-binding proteins, RNA modifications and additional factors in nuclear and cytoplasmic processes, including alternative splicing, 3' end processing, nuclear surveillance, and RNA decay.
Carmen Aguilar

Carmen Aguilar © Hilde Merkert
Host Pathways in Urinary Tract Infections
Urinary tract infections (UTIs) caused by Gram-negative uropathogenic Escherichia coli (UPEC) are among the most common and recurrent infections worldwide. Up to now, most of the anti-infective research has been focused on combating the bacterial side. However, whether a person is susceptible to infection by UPEC is also decided by host factors. Hence, host-based therapeutics represent an innovative approach to fight infections. The Aguilar group aims to obtain a better understanding of the host pathways controlling UPEC infection and persistence in the bladder. The researchers use adult stem cell derived-organoids and organoid-based models to closely mimic the natural infection site. They combine these infection models with miRNome profiling (global analysis of host microRNA expression), single-cell RNA-sequencing, and gain/loss-of-function approaches in order to identify and functionally characterize specific host factors and/or pathways that are required for UPEC to infect and persist in the human bladder epithelium. Based on this knowledge, new host-based therapeutic approaches could be designed to prevent or treat UTIs.
Simone Backes

Simone Backes © privat
Roles of RNAs During Virus Infection
The Backes group is interested in studying RNA-based mechanisms in virulence, virus replication and host defense.
Irene Beusch

Irene Beusch © IMIB
Regulation and Mechanisms in RNA Splicing
The Beusch group studies regulatory mechanisms which steer the assembly of the spliceosome but also allow it to handle its very diverse substrate pool. We tackle this problem with an interdisciplinary approach by combining genetics & genomics with molecular biology & biochemistry. By gaining a detailed mechanistic understanding of splicing and its regulation within the cell, we can not only understand fundamental eukaryotic biology but also how its dysregulation impacts human disease.
Dimitrios Papadopoulos

Dimitrios Papadopoulos © privat
RNA-based Mechanisms of Tumor Maintenance
The Papadopoulos group focuses on understanding the molecular mechanisms that maintain aggressive human cancers. We are particularly interested in the MYC oncoprotein family and how their newly discovered RNA-binding functions mitigate DNA damage and prevent immune recognition in rapidly proliferating cancer cells.
Karl Petri

Karl Petri © JMU / Daniel Peter
Innovating RNA-guided Genome Engineering for Cancer-Directed Cellular Immunotherapy
The goal of the Petri group is to deploy novel RNA-guided gene editing technology to enhance immune cell therapies. To do this, we a) develop new RNA-guided editing tools, b) evaluate their safety using state-of-the-art molecular methods, and c) apply them to enhance immune cell therapies. With a particular focus on oncologic diseases, we aim to streamline the application and safety-validation of RNA-guided gene editing tools to generate and enhance immune cell products for cancer patients.

More about RNA research in Würzburg
The research foci of the Helmholtz Institute























