Prof Lars Barquist, Associated Scientist
Integrative Informatics for Infection Biology
In 2018, the Helmholtz Institute Würzburg (HIRI) established a research group for integrative informatics in infection biology. Group leader Lars Barquist, who has been appointed assistant professor at the University of Toronto in Canada in 2024, coordinates and advises the group with the aim to develop new statistical, bioinformatic, and visualization approaches in infection biology.
Our research and approach
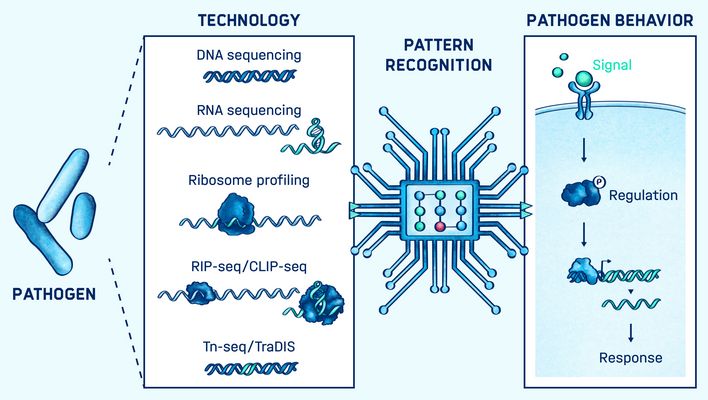
The development of high-throughput technologies in infection biology allows thousands of genetic loci to be interrogated simultaneously in a single experiment. This has resulted in an accumulation of gene expression, fitness, and regulatory interaction data. The bottleneck in advancing our understanding of pathogens now lies in moving from hypothesis-free screening through data integration to hypothesis generation.
Lars Barquist's group works on the development of new statistical, bioinformatic, and visualization approaches. Their work applies tools such as machine learning and Bayesian statistics to provide insight into bacterial pathogens, with the ultimate aim of overcoming the current bottleneck in the interpretation of complex post-genomic data.
Together with other research groups at the HIRI, the group has worked on the application of a variety of technologies, including CLIP seq, Dual RNA seq, CRISPRi screens, transposon mutagenesis and meta-transcriptomics. Their overarching goal is to bring together sequencing data and new computational technologies to advance our understanding of bacterial pathogenesis and the development of RNA-based therapeutics.
Team members

Prof Lars Barquist, Associated Scientist
Group Leader

Dr Laura Jenniches
Postdoc

Dr Willow Kion-Crosby
Visiting Scientist
Research projects
Graphical Abstract
Alumni
Shuba Varshini Alampalli, PhD Student • Regan Hayward, Postdoc • Rituparno Sen, Postdoc • Eva Weiß, Postdoc • Yanying Yu, PhD Student
Publications
2026
Stress-induced iron-sulfur cluster damage as a conserved trigger of the bacterial stringent response
Michaud E, Ricci L, Lallement C, Barquist L, Vincent C, Michaux C, Hallez R, Ronneau S, Michaud E, Ricci L, …, Hallez R, Ronneau S (2026)
Nature Communications 17 (1)
2025
High-Throughput Tiling of Essential mRNAs Increases Potency of Antisense Antibiotics
Danti G, Popella L, Vogel J, Maric HM (2025)
Advanced Science 12 (28): e2504284
The global RNA-binding protein RbpB is a regulator of polysaccharide utilization in Bacteroides thetaiotaomicron
Rüttiger AS, Ryan D, Spiga L, Lamm-Schmidt V, Prezza G, Reichardt S, Langford M, Barquist L, Faber F, Zhu W, Westermann AJ (2025)
Nature Communications 16 (1): 208
Biased sampling driven by bacterial population structure confounds machine learning prediction of antimicrobial resistance
Yu Y, Wheeler NE, Barquist L (2025)
PLOS Biology 23 (12): e3003539
The pcnB gene sustains Shigella flexneri virulence
Frisch T, Geiser P, Komi M, Karlsson PA, Bhetwal A, Jenniches L, Barquist L, Holmqvist E, Mateus A, Sellin ME, Di Martino ML (2025)
PLOS Pathogens 21 (11): e1013727
Low-input RNA-seq suggests metabolic specialization underlying morphological heterogeneity in a gut commensal bacterium
Bornet E, Prezza G, Cecchino L, Jenniches L, Behrends J, Tawk C, Huang KC, Strowig T, Vogel J, Barquist L, Saliba AE, Westermann AJ (2025)
Cell Reports 44 (6): 115844
A scalable gut epithelial organoid model reveals the genome-wide colonization landscape of a human-adapted pathogen
Di Martino ML, Jenniches L, Bhetwal A, Eriksson J, Lopes ACC, Ntokaki A, Pasqua M, Sundbom M, Skogar M, Graf W, …, Barquist L, Sellin ME (2025)
Nature Genetics 57 (7): 1730-1741
A Small RNA Derived From the 5' End of the IS200 tnpA Transcript Regulates Multiple Virulence Regulons in Salmonella typhimurium
Trussler RS, Scherba NQ, Kooshapour H, Ellis MJ, Förstner KU, Albert M, Westermann AJ, Haniford DB (2025)
Molecular Microbiology 124 (5): 413-432
An antisense oligomer conjugate with unpredicted bactericidal activity against Fusobacterium nucleatum
Cosi V, Jung J, Popella L, Ponath F, Ghosh C, Barquist L, Vogel J (2025)
mBio 16 (6): e0052425
Micromix: web infrastructure for visualizing and remixing microbial 'omics data
Hayward RJ, Ebbecke T, Fricke H, Nguyen VQ, Barquist L (2025)
GigaScience 14: giae120
2024
sCIRCLE-An interactive visual exploration tool for single cell RNA-Seq data
Seeger M, Schöls E, Barquist L (2024)
NAR Genomics and Bioinformatics 6 (3): lqae084
A cell-free transcription-translation pipeline for recreating methylation patterns boosts DNA transformation in bacteria
Vento JM, Durmusoglu D, Li T, Patinios C, Sullivan S, Ttofali F, van Schaik J, Yu Y, Wang Y, Barquist L, Crook N, Beisel CL (2024)
Molecular Cell 84 (14): 2785-2796.e4
Expanding the flexibility of base editing for high-throughput genetic screens in bacteria
Gawlitt S, Collins SP, Yu Y, Blackman SA, Barquist L, Beisel CL (2024)
Nucleic Acids Research 52 (7): 4079-4097
Improved RNA stability estimation through Bayesian modeling reveals most Salmonella transcripts have subminute half-lives
Jenniches L, Michaux C, Popella L, Reichardt S, Vogel J, Westermann AJ, Barquist L (2024)
PNAS 121 (14): e2308814121
An expanded transcriptome atlas for Bacteroides thetaiotaomicron reveals a small RNA that modulates tetracycline sensitivity
Ryan D, Bornet E, Prezza G, Alampalli SV, Franco de Carvalho T, Felchle H, Ebbecke T, Hayward RJ, Deutschbauer AM, Barquist L, Westermann AJ (2024)
Nature Microbiology 9 (4): 1130-1144
Improved prediction of bacterial CRISPRi guide efficiency from depletion screens through mixed-effect machine learning and data integration
Yu Y, Gawlitt S, de Andrade E Sousa LB, Merdivan E, Piraud M, Beisel CL, Barquist L (2024)
Genome Biology 25 (1): 13
The conserved noncoding RNA ModT coordinates growth and virulence in Clostridioides difficile
Lence T, Sulzer J, Andress K, Gribling-Burrer AS, Lamm-Schmidt V, Barquist L, Smyth RP, Faber F (2024)
PLOS Biology 22 (12): e3002948
Network depth affects inference of gene sets from bacterial transcriptomes using denoising autoencoders
Kion-Crosby W, Barquist L (2024)
Bioinformatics Advances 4 (1): 181
A method to correct for local alterations in DNA copy number that bias functional genomics assays applied to antibiotic-treated bacteria
Sullivan GJ, Barquist L, Cain AK (2024)
mSystems 12: e0066523
2023
Vector-borne Trypanosoma brucei parasites develop in artificial human skin and persist as skin tissue forms
Reuter C, Hauf L, Imdahl F, Sen R, Vafadarnejad E, Fey P, Finger T, Jones NG, Walles H, Barquist L, …, Groeber-Becker F, Engstler M (2023)
Nature Communications 14 (1): 7660
Ribosome profiling reveals the fine-tuned response of Escherichia coli to mild and severe acid stress
Schumacher K, Gelhausen R, Kion-Crosby W, Barquist L, Backofen R, Jung K (2023)
mSystems 8 (6): e0103723
Grad-seq analysis of Enterococcus faecalis and Enterococcus faecium provides a global view of RNA and protein complexes in these two opportunistic pathogens
Michaux C, Gerovac M, Hansen EE, Barquist L, Vogel J (2023)
microLife 4: uqac027
DksA is a conserved master regulator of stress response in Acinetobacter baumannii
Maharjan RP, Sullivan GJ, Adams FG, Shah BS, Hawkey J, Delgado N, Semenec L, Dinh H, Li L, Short FL, …, Eijkelkamp BA, Cain AK (2023)
Nucleic Acids Research 51 (12): 6101-6119
Improved Bacterial Single-Cell RNA-Seq through Automated MATQ-Seq and Cas9-Based Removal of rRNA Reads
Homberger C, Hayward RJ, Barquist L, Vogel J (2023)
mBio 14 (2): e0355722
Design and off-target prediction for antisense oligomers targeting bacterial mRNAs with the MASON webserver
Jung JJ, Popella L, Do PT, Pfau P, Vogel J, Barquist L (2023)
RNA 299 (5): 570-583
2022
Network community detection and clustering with random walks
Ballal A, Kion-Crosby W, Morozov A (2022)
Physical Review Research 4 (4): 583
A target expression threshold dictates invader defense and prevents autoimmunity by CRISPR-Cas13
Vialetto E, Yu Y, Collins SP, Wandera KG, Barquist L, Beisel CL (2022)
Cell Host & Microbe 3128 (22): 00273-6
Comprehensive analysis of PNA-based antisense antibiotics targeting various essential genes in uropathogenic Escherichia coli
Popella L, Jung J, Do PT, Hayward RJ, Barquist L, Vogel J (2022)
Nucleic Acids Research 50 (11): 6435–6452
Ushering in a new era of single-cell transcriptomics in bacteria
Homberger C, Barquist L, Vogel J (2022)
microLife 3: uqac020
Cellular RNA Targets of Cold Shock Proteins CspC and CspE and Their Importance for Serum Resistance in Septicemic Escherichia coli
Yair Y, Michaux C, Biran D, Bernhard J, Vogel J, Barquist L, Ron EZ (2022)
mSystems 7 (4): e0008622
Comparative genomics provides structural and functional insights into Bacteroides RNA biology
Prezza G, Ryan D, Mädler G, Reichardt S, Barquist L, Westermann AJ (2022)
Molecular Microbiology 117 (1): 67-85
2021
RNA landscape of the emerging cancer-associated microbe Fusobacterium nucleatum
Ponath F, Tawk C, Zhu Y, Barquist L, Faber F, Vogel J (2021)
Nature Microbiology 6 (8): 1007-1020
Noncanonical crRNAs derived from host transcripts enable multiplexable RNA detection by Cas9
Jiao C, Sharma S, Dugar G, Peeck NL, Bischler T, Wimmer F, Yu Y, Barquist L, Schoen C, Kurzai O, Sharma CM, Beisel CL (2021)
Science 372 (6545): 941-948
Global RNA profiles show target selectivity and physiological effects of peptide-delivered antisense antibiotics
Popella L, Jung J, Popova K, Ðurica-Mitic S, Barquist L, Vogel J (2021)
Nucleic Acids Research 49 (8): 4705-4724
Common Features in lncRNA Annotation and Classification: A Survey
Klapproth C, Sen R, Stadler PF, Findeiß S, Fallmann J (2021)
Non-Coding RNA 7 (4): 77
An RNA-centric global view of Clostridioides difficile reveals broad activity of Hfq in a clinically important gram-positive bacterium
Fuchs M, Lamm-Schmidt V, Sulzer J, Ponath F, Jenniches L, Kirk JA, Fagan RP, Barquist L, Vogel J, Faber F (2021)
PNAS 118 (25): e2103579118
Functional analysis of colonization factor antigen I positive enterotoxigenic Escherichia coli identifies genes implicated in survival in water and host colonization
Abd El Ghany M, Barquist L, Clare S, Brandt C, Mayho M, Joffre E, Sjöling Å, Turner AK, Klena JD, Kingsley RA, …, Dougan G, Pickard D (2021)
Microbial Genomics 7 (6): 000554
Global identification of RsmA/N binding sites in Pseudomonas aeruginosa by in vivo UV CLIP-seq
Chihara K, Barquist L, Takasugi K, Noda N, Tsuneda S (2021)
RNA Biology 18 (12): 2401-2416
Global identification of RsmA/N binding sites in Pseudomonas aeruginosa by in vivo UV CLIP-seq
Chihara K, Barquist L, Takasugi K, Noda N, Tsuneda S (2021)
RNA Biology 18 (12): 2401-2416
2020
A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron
Ryan D, Jenniches L, Reichardt S, Barquist L, Westermann AJ (2020)
Nature Communications 11 (1): 3557
Dual RNA-seq of Orientia tsutsugamushi informs on host-pathogen interactions for this neglected intracellular human pathogen
Mika-Gospodorz B, Giengkam S, Westermann AJ, Wongsantichon J, Kion-Crosby W, Chuenklin S, Wang LC, Sunyakumthorn P, Sobota RM, Subbian S, …, Barquist L, Salje J (2020)
Nature Communications 11: 3363
An amphipathic peptide with antibiotic activity against multidrug-resistant Gram-negative bacteria
Elliott AG, Huang JX, Neve S, Zuegg J, Edwards IA, Cain AK, Boinett CJ, Barquist L, Lundberg CV, Steen J, …, Strandh M, Cooper MA (2020)
Nature Communications 11: 3184
A decade of advances in transposon-insertion sequencing
Cain AK, Barquist L, Goodman AL, Paulsen IT, Parkhill J, van Opijnen T (2020)
Nature Reviews Genetics 21 (9): 526-540
The minimal meningococcal ProQ protein has an intrinsic capacity for structure-based global RNA recognition
Bauriedl S, Gerovac M, Heidrich N, Bischler T, Barquist L, Vogel J, Schoen C (2020)
Nature Communications 11: 2823
A global data-driven census of Salmonella small proteins and their potential functions in bacterial virulence
Venturini E, Svensson SL, Maaß S, Gelhausen R, Eggenhofer F, Li L, Cain AK, Parkhill J, Becher D, Backofen R, …, Westermann AJ, Vogel J (2020)
microLife 1 (1): 597
Single-Nucleotide RNA Maps for the Two Major Nosocomial Pathogens Enterococcus faecalis and Enterococcus faecium
Michaux C, Hansen EE, Jenniches L, Gerovac M, Barquist L, Vogel J (2020)
Frontiers in Cellular and Infection Microbiology 10: 600325
Whole genome sequencing of Thametopoea pityocampa revealed putative pesticide targets
Shahraki A, Yu Y, Gul ZM, Liang C, Iyison NB (2020)
Genomics 112 (6): 4203-4207
Global discovery of bacterial RNA-binding proteins by RNase-sensitive gradient profiles reports a new FinO domain protein
Gerovac M, El Mouali Y, Kuper J, Kisker C, Barquist L, Vogel J (2020)
RNA 26 (10): 1448-1463
2019
Conditional Hfq Association with Small Noncoding RNAs in Pseudomonas aeruginosa Revealed through Comparative UV Cross-Linking Immunoprecipitation Followed by High-Throughput Sequencing
Chihara K, Bischler T, Barquist L, Monzon VA, Noda N, Vogel J, Tsuneda S (2019)
mSystems 4 (6): e00590-19
Rapid transcriptional responses to serum exposure are associated with sensitivity and resistance to antibody-mediated complement killing in invasive Salmonella Typhimurium ST313
Ondari EM, Klemm EJ, Msefula CL, El Ghany MA, Heath JN, Pickard DJ, Barquist L, Dougan G, Kingsley RA, MacLennan CA (2019)
Wellcome Open Research 4: 74
Transcriptional noise and exaptation as sources for bacterial sRNAs
Jose BR, Gardner PP, Barquist L (2019)
Biochemical Society Transactions 47 (2): 527-539
2018
Global Maps of ProQ Binding In Vivo Reveal Target Recognition via RNA Structure and Stability Control at mRNA 3' Ends
Holmqvist E, Li L, Bischler T, Barquist L, Vogel J (2018)
Molecular Cell 70 (5): 971-982.e6
A global genomic approach uncovers novel components for twitching motility-mediated biofilm expansion in Pseudomonas aeruginosa
Nolan LM, Whitchurch CB, Barquist L, Katrib M, Boinett CJ, Mayho M, Goulding D, Charles IG, Filloux A, Parkhill J, Cain AK (2018)
Microbial Genomics 4 (11): e000229
Functional analysis of Salmonella Typhi adaptation to survival in water
Kingsley RA, Langridge G, Smith SE, Makendi C, Fookes M, Wileman TM, El Ghany MA, Keith Turner A, Dyson ZA, Sridhar S, …, Barquist L, Dougan G (2018)
Environmental Microbiology 20 (11): 4079-4090
Morphological, genomic and transcriptomic responses of Klebsiella pneumoniae to the last-line antibiotic colistin
Cain AK, Boinett CJ, Barquist L, Dordel J, Fookes M, Mayho M, Ellington MJ, Goulding D, Pickard D, Wick RR, …, Parkhill J, Thomson NR (2018)
Scientific Reports 8 (1): 9868
Machine learning identifies signatures of host adaptation in the bacterial pathogen Salmonella enterica
Wheeler NE, Gardner PP, Barquist L (2018)
PLOS Genetics 14 (5): e1007333
2017
The primary transcriptome of Neisseria meningitidis and its interaction with the RNA chaperone Hfq
Heidrich N, Bauriedl S, Barquist L, Li L, Schoen C, Vogel J (2017)
Nucleic Acids Research 45 (10): 6147-6167
RNA target profiles direct the discovery of virulence functions for the cold-shock proteins CspC and CspE
Michaux C, Holmqvist E, Vasicek E, Sharan M, Barquist L, Westermann AJ, Gunn JS, Vogel J (2017)
PNAS 114 (26): 6824-6829
Role of sapA and yfgA in Susceptibility to Antibody-Mediated Complement-Dependent Killing and Virulence of Salmonella enterica Serovar Typhimurium
Ondari EM, Heath JN, Klemm EJ, Langridge G, Barquist L, Goulding DA, Clare S, Dougan G, Kingsley RA, MacLennan CA (2017)
Infection and Immunity 85 (9): pii: e00419-17
Resolving host-pathogen interactions by dual RNA-seq
Westermann AJ, Barquist L, Vogel J (2017)
PLOS Pathogens 13 (2): e1006033
2016
Distinct Salmonella Enteritidis lineages associated with enterocolitis in high-income settings and invasive disease in low-income settings
Feasey NA, Hadfield J, Keddy KH, Dallman TJ, Jacobs J, Deng X, Wigley P, Barquist L, Langridge GC, Feltwell T, …, Gordon MA, Thomson NR (2016)
Nature Genetics 48 (10): 1211-1217
Dual RNA-seq unveils noncoding RNA functions in host-pathogen interactions
Westermann AJ, Forstner KU, Amman F, Barquist L, Chao Y, Schulte LN, Muller L, Reinhardt R, Stadler PF, Vogel J (2016)
Nature 529 (7587): 496-501
Global RNA recognition patterns of post-transcriptional regulators Hfq and CsrA revealed by UV crosslinking in vivo
Holmqvist E, Wright PR, Li L, Bischler T, Barquist L, Reinhardt R, Backofen R, Vogel J (2016)
The EMBO Journal 35 (9): 991-1011
The TraDIS toolkit: sequencing and analysis for dense transposon mutant libraries
Barquist L, Mayho M, Cummins C, Cain AK, Boinett CJ, Page AJ, Langridge GC, Quail MA, Keane JA, Parkhill J (2016)
Bioinformatics 32 (7): 1109-11
Studying RNA Homology and Conservation with Infernal: From Single Sequences to RNA Families
Barquist L, Burge SW, Gardner PP (2016)
Current Protocols in Bioinformatics 54: 12.13.1-12.13.25
Molecular phenotyping of infection-associated small non-coding RNAs
Barquist L, Westermann AJ, Vogel J (2016)
Philosophical Transactions of the Royal Society of London B: Biological Sciences 371 (1707): 20160081
The in vitro and in vivo effects of constitutive light expression on a bioluminescent strain of the mouse enteropathogen Citrobacter rodentium
Read HM, Mills G, Johnson S, Tsai P, Dalton J, Barquist L, Print CG, Patrick WM, Wiles S (2016)
PeerJ 4: e2130
A profile-based method for identifying functional divergence of orthologous genes in bacterial genomes
Wheeler NE, Barquist L, Kingsley RA, Gardner PP (2016)
Bioinformatics 32 (23): 3566-3574
2015
Accelerating Discovery and Functional Analysis of Small RNAs with New Technologies
Barquist L, Vogel J (2015)
Annual Review of Genetics 49: 367-94
High-throughput analysis of gene essentiality and sporulation in Clostridium difficile
Dembek M, Barquist L, Boinett CJ, Cain AK, Mayho M, Lawley TD, Fairweather NF, Fagan RP (2015)
mBio 6 (2): e02383
Patterns of genome evolution that have accompanied host adaptation in Salmonella
Langridge GC, Fookes M, Connor TR, Feltwell T, Feasey N, Parsons BN, Seth-Smith HM, Barquist L, Stedman A, Humphrey T, …, Dougan G, Thomson NR (2015)
PNAS 112 (3): 863-8
Signatures of adaptation in human invasive Salmonella Typhimurium ST313 populations from sub-Saharan Africa
Okoro CK, Barquist L, Connor TR, Harris SR, Clare S, Stevens MP, Arends MJ, Hale C, Kane L, Pickard DJ, …, Dougan G, Kingsley RA (2015)
PLOS Neglected Tropical Diseases 9 (3): e0003611
2014
Functional genomics reveals that Clostridium difficile Spo0A coordinates sporulation, virulence and metabolism
Pettit LJ, Browne HP, Yu L, Smits WK, Fagan RP, Barquist L, Martin MJ, Goulding D, Duncan SH, Flint HJ, …, Choudhary JS, Lawley TD (2014)
BMC Genomics 15: 160
Robust identification of noncoding RNA from transcriptomes requires phylogenetically-informed sampling
Lindgreen S, Umu SU, Lai AS, Eldai H, Liu W, McGimpsey S, Wheeler NE, Biggs PJ, Thomson NR, Barquist L, Poole AM, Gardner PP (2014)
PLOS Computational Biology 10 (10): e1003907
Parallel independent evolution of pathogenicity within the genus Yersinia
Reuter S, Connor TR, Barquist L, Walker D, Feltwell T, Harris SR, Fookes M, Hall ME, Petty NK, Fuchs TM, …, McNally A, Thomson NR (2014)
PNAS 111 (18): 6768-73
An introduction to RNA databases
Hoeppner MP, Barquist L, Gardner PP (2014)
In: RNA Sequence, Structure, and Function: Computational and Bioinformatic Methods (eds Gorodkin J, Ruzzo W), Methods Mol Biol 1097: 107-23
2013
Rfam 11.0: 10 years of RNA families
Burge SW, Daub J, Eberhardt R, Tate J, Barquist L, Nawrocki EP, Eddy SR, Gardner PP, Bateman A (2013)
Nucleic Acids Research 41 (Database issue): D226-32
RNA-seq reveals the RNA binding proteins, Hfq and RsmA, play various roles in virulence, antibiotic production and genomic flux in Serratia sp. ATCC 39006
Wilf NM, Reid AJ, Ramsay JP, Williamson NR, Croucher NJ, Gatto L, Hester SS, Goulding D, Barquist L, Lilley KS, …, Dougan G, Salmond GP (2013)
BMC Genomics 14: 822
A comparison of dense transposon insertion libraries in the Salmonella serovars Typhi and Typhimurium
Barquist L, Langridge GC, Turner DJ, Phan M, Turner AK, Bateman A, Parkhill J, Wain J, Gardner PP (2013)
Nucleic Acids Research 41 (8): 4549-64
Dominant role of nucleotide substitution in the diversification of serotype 3 pneumococci over decades and during a single infection
Croucher NJ, Mitchell AM, Gould KA, Inverarity D, Barquist L, Feltwell T, Fookes MC, Harris SR, Dordel J, Salter SJ, …, Mitchell TJ, Bentley SD (2013)
PLOS Genetics 9 (10): e1003868
The agr locus regulates virulence and colonization genes in Clostridium difficile 027
Martin MJ, Clare S, Goulding D, Faulds-Pain A, Barquist L, Browne HP, Pettit LJ, Dougan G, Lawley TD, Wren BW (2013)
Journal of Bacteriology 195 (16): 3672-81
Approaches to querying bacterial genomes with transposon-insertion sequencing
Barquist L, Boinett CJ, Cain AK (2013)
RNA Biology 10 (7): 1161-9
Genome and transcriptome adaptation accompanying emergence of the definitive type 2 host-restricted Salmonella enterica serovar Typhimurium pathovar
Kingsley RA, Kay S, Connor T, Barquist L, Sait L, Holt KE, Sivaraman K, Wileman TM, Goulding D, Clare S, …, Parkhill J, Dougan G (2013)
mBio 4 (5): e00565-13
Characterization of the yehUT two-component regulatory system of Salmonella enterica Serovar Typhi and Typhimurium
Wong VK, Pickard DJ, Barquist L, Sivaraman K, Page AJ, Hart PJ, Arends MJ, Holt KE, Kane L, Mottram LF, …, Kingsley RA, Dougan G (2013)
PLOS One 8 (12): e84567
2012
A high-resolution view of genome-wide pneumococcal transformation
Croucher NJ, Harris SR, Barquist L, Parkhill J, Bentley SD (2012)
PLOS Pathogens 8 (6): e1002745
HandAlign: Bayesian multiple sequence alignment, phylogeny and ancestral reconstruction
Westesson O, Barquist L, Holmes I (2012)
Bioinformatics 28 (8): 1170-1
2011
RNIE: genome-wide prediction of bacterial intrinsic terminators
Gardner PP, Barquist L, Bateman A, Nawrocki EP, Weinberg Z (2011)
Nucleic Acids Research 39 (14): 5845-52
2009
Evolutionary modeling and prediction of non-coding RNAs in Drosophila
Bradley RK, Uzilov AV, Skinner ME, Bendana YR, Barquist L, Holmes I (2009)
PLOS One 4 (8): e6478
2008
xREI: a phylo-grammar visualization webserver
Barquist L, Holmes I (2008)
Nucleic Acids Research 36 (Web Server issue): W65-W69