Prof Neva Caliskan, Former Group Leader
Recoding Mechanisms in Infections
In 2018, the Helmholtz Institute Würzburg (HIRI) established a research group on recoding mechanisms in infection, headed by Neva Caliskan. While maintaining close ties to research at the HIRI, Caliskan was appointed professor and head of the Department of Biochemistry III at the University of Regensburg in 2024.
Our research and approach
Viruses and cellular genes encode RNAs that can be read in alternative ways during translation, which is called recoding. However, how exactly recoding is regulated by host encoded factors remains elusive. Here, a detailed understanding of recoding and its regulation can open doors for the development of novel RNA-based therapeutic interventions to combat infections.
Neva Caliskan's group investigated the functions and dynamics of RNA molecules and their interplay with trans-acting factors involved in recoding events. They worked with several viruses known to depend on recoding strategies for replication including corona and retroviruses, and developed methods to investigate RNA complexes and translation in unprecedented detail.
The group employed a highly interdisciplinary toolset including RNA-antisense purification and mass spectrometry to identify RNA-interaction partners, and cellular assays to investigate molecular details. Ensemble and single molecule assays such as optical tweezers are key to study the dynamics of RNA complexes. Ultimately, they sought to understand how RNA-structure elements act in concert with other factors in the cell to modulate the way mRNA messages are read by ribosomes during infections to advance RNA-based therapeutics.
Team members

Prof Neva Caliskan, Former Group Leader
Group Leader
Research projects
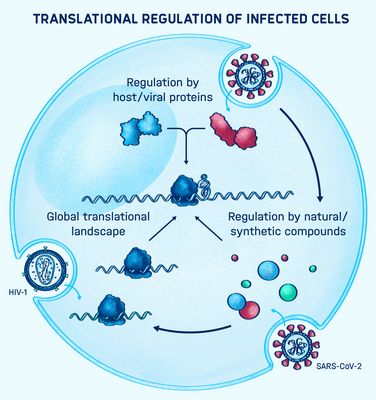
Graphical Abstract
Alumni
Stefan Buck, Data Manager • Orian Gilmer, Postdoc • Anuja Kibe, PhD Student • Tatyana Koch, Technical Assistant • Saurav Malik, PhD Student • Rui Pedro Marques da Costa, Postdoc • Lukáš Pekárek, PhD Student • Ricarda Riegger, PhD Student • Alexandria Smart, Postdoc • Matthias Zimmer, PhD Student & Postdoc
Publications
2025
Guidelines for minimal reporting requirements, design and interpretation of experiments involving the use of eukaryotic dual gene expression reporters (MINDR)
Loughran G, Andreev DE, Terenin IM, Namy O, Mikl M, Yordanova MM, McManus CJ, Firth AE, Atkins JF, Fraser CS, …, Dmitriev SE, Baranov PV (2025)
Nature Structural & Molecular Biology 32 (3): 418-430
Optimization of Structure-Guided Development of Chemical Probes for the Pseudoknot RNA of the Frameshift Element in SARS-CoV-2
Ceylan B, Adam J, Toews S, Kaiser F, Dörr J, Scheppa D, Tants JN, Smart A, Schoth J, Philipp S, …, Wacker A, Schwalbe H (2025)
Angewandte Chemie - International Edition 64 (9): e202417961
AcrVIB1 inhibits CRISPR-Cas13b immunity by promoting unproductive crRNA binding accessible to RNase attack
Wandera KG, Schmelz S, Migur A, Kibe A, Lukat P, Achmedov T, Caliskan N, Blankenfeldt W, Beisel CL (2025)
Molecular Cell 85 (6): 1162-1175
The translational landscape of HIV-1 infected cells reveals key gene regulatory principles
Kibe A, Buck S, Gribling-Burrer AS, Gilmer O, Bohn P, Koch T, Mireisz CN, Schlosser A, Erhard F, Smyth RP, Caliskan N (2025)
Nature Structural & Molecular Biology 32 (5): 841-852
High-Fidelity and Cost-Effective Engineering of SARS-CoV-2
Olguin-Nava M, Hennig T, Börtlein C, Bohn P, Ambi UB, Gabel A, Günter LM, Gribling-Burrer AS, Schmidt N, Caliskan N, …, Munschauer M, Smyth RP (2025)
Viruses 17 (12): 1604
Type I-Fv and engineered type IV-A1 CRISPR-Cas effectors facilitate genome reduction in Escherichia coli
Klein N, Sanchez-Londono M, Kara MM, Gomes-Filho JV, Novak S, Kholeif KH, Pekarek L, Caliskan N, Randau L (2025)
Nucleic Acids Research 53 (22): gkaf1399
Designing the Aplysia punctata Arginine-Depleting Enzyme for Tumor Targeting
Wolkersdorfer AM, Endo Y, Kehrein J, Kappus M, Hattori S, Gutmann M, Rudel T, Caliskan N, Lühmann T, Kato Y, Meinel L (2025)
Molecular Pharmaceutics 22 (3): 1253–1261
Giant RNA genomes: Roles of host, translation elongation, genome architecture, and proteome in nidoviruses
Neuman BW, Smart A, Gilmer O, Smyth RP, Vaas J, Böker N, Samborskiy DV, Bartenschlager R, Seitz S, Gorbalenya AE, Caliskan N, Lauber C (2025)
PNAS 122 (7): e2413675122
2024
Translation Inhibition Mediated by Interferon-Stimulated Genes during Viral Infections
Smart A, Gilmer O, Caliskan N (2024)
Viruses 16 (7)
2023
Cis-mediated interactions of the SARS-CoV-2 frameshift RNA alter its conformations and affect function
Pekarek L, Zimmer MM, Gribling-Burrer AS, Buck S, Smyth RP, Caliskan N (2023)
Nucleic Acids Research 51 (2): 728–743
SND1 binds SARS-CoV-2 negative-sense RNA and promotes viral RNA synthesis through NSP9
Schmidt N, Ganskih S, Wei Y, Gabel A, Zielinski S, Keshishian H, Lareau CA, Zimmermann L, Makroczyova J, Pearce C, …, Erhard F, Munschauer M (2023)
Cell 186 (22): 4834-4850.e23
Mouse Liver-Expressed Shiftless Is an Evolutionarily Conserved Antiviral Effector Restricting Human and Murine Hepaciviruses
Zhang Y, Kinast V, Sheldon J, Frericks N, Todt D, Zimmer M, Caliskan N, Brown RJP, Steinmann E, Pietschmann T (2023)
Microbiology Spectrum 11 (4): e0128423
2022
Short- and long-range interactions in the HIV-1 5' UTR regulate genome dimerization and packaging
Ye L, Gribling-Burrer AS, Bohn P, Kibe A, Börtlein C, Ambi UB, Ahmad S, Olguin-Nava M, Smith M, Caliskan N, von Kleist M, Smyth RP (2022)
Nature Structural & Molecular Biology 29 (4): 306-319
Spacer prioritization in CRISPR-Cas9 immunity is enabled by the leader RNA
Liao C, Sharma S, Svensson SL, Kibe A, Weinberg Z, Alkhnbashi OS, Bischler T, Backofen R, Caliskan N, Sharma CM, Beisel CL (2022)
Nature Microbiology 7 (4): 530-541
Editorial: mRNA Translational Control as a Mechanism of Post-transcriptional Gene Regulation
Kiss DL, Vasudevan D, Ho CK, Caliskan N (2022)
Frontiers in Molecular Biosciences 9: 947516
POTATO: Automated pipeline for batch analysis of optical tweezers data
Buck S, Pekarek L, Caliskan N (2022)
Biophysical Journal 121 (15): 2830-2839
Insights from structural studies of the cardiovirus 2A protein
Caliskan N, Hill CH (2022)
Bioscience Reports 42 (1): BSR20210406
Optical Tweezers to Study RNA-Protein Interactions in Translation Regulation
Pekarek L, Buck S, Caliskan N (2022)
Journal of Visualized Experiments (180)
Thinking Outside the Frame: Impacting Genomes Capacity by Programmed Ribosomal Frameshifting
Riegger RJ, Caliskan N (2022)
Frontiers in Molecular Biosciences 9: 842261
2021
Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Hill CH, Pekarek L, Napthine S, Kibe A, Firth AE, Graham SC, Caliskan N, Brierley I (2021)
Nature Communications 12 (1): 7166
Investigating molecular mechanisms of 2A-stimulated ribosomal pausing and frameshifting in Theilovirus
Hill CH, Cook GM, Napthine S, Kibe A, Brown K, Caliskan N, Firth AE, Graham SC, Brierley I (2021)
Nucleic Acids Research 49 (20): 11938-11958
The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting
Zimmer MM, Kibe A, Rand U, Pekarek L, Ye L, Buck S, Smyth RP, Cicin-Sain L, Caliskan N (2021)
Nature Communications 12 (1): 7193
2020
The SARS-CoV-2 RNA-protein interactome in infected human cells
Schmidt N, Lareau CA, Keshishian H, Ganskih S, Schneider C, Hennig T, Melanson R, Werner S, Wei Y, Zimmer M, …, Bodem J, Munschauer M (2020)
Nature Microbiology 6 (3): 339-353
2019
Thermodynamic control of -1 programmed ribosomal frameshifting
Bock LV, Caliskan N, Korniy N, Peske F, Rodnina MV, Grubmüller H (2019)
Nature Communications 10: 4598
2018
Small synthetic molecule-stabilized RNA pseudoknot as an activator for -1 ribosomal frameshifting
Matsumoto S, Caliskan N, Rodnina MV, Murata A, Nakatani K (2018)
Nucleic Acids Research 46 (16): 8079-8089
2017
Conditional Switch between Frameshifting Regimes upon Translation of dnaX mRNA
Caliskan N, Wohlgemuth I, Korniy N, Pearson M, Peske F, Rodnina MV (2017)
Molecular Cell 66 (4): 558-567.e4
2016
Choreography of molecular movements during ribosome progression along mRNA
Belardinelli R, Sharma H, Caliskan N, Cunha CE, Peske F, Wintermeyer W, Rodnina MV (2016)
Nature Structural & Molecular Biology 23 (4): 342-8
2015
Changed in translation: mRNA recoding by -1 programmed ribosomal frameshifting
Caliskan N, Peske F, Rodnina MV (2015)
Trends in Biochemical Sciences 40 (5): 265-74
2014
Programmed -1 frameshifting by kinetic partitioning during impeded translocation
Caliskan N, Katunin VI, Belardinelli R, Peske F, Rodnina MV (2014)
Cell 157 (7): 1619-31