Prof Alexander Westermann, Affiliated Scientist
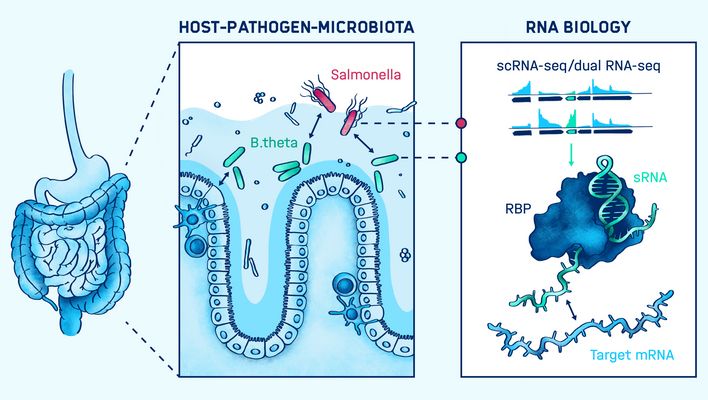
Host-pathogen-microbiota interactions
Interactions between the host, invading pathogens and the host's gut microbiota have been investigated at the HIRI since 2018. From 2018 to 2023, Alexander Westermann headed a correspondent research group. Since 2023, Westermann holds a professorship at the Chair of Microbiology at the University of Würzburg and contributes to the research at the HIRI as an affiliated scientist.
Our research and approach
The current antibiotics crisis has highlighted the urgency of developing alternative therapeutic approaches to infectious disease. RNA-based therapies offer great potential for targeting specific enteric pathogens at species level, whilst minimizing collateral damage to the surrounding microbiota.
Alexander Westermann's group mapped the complex network of interactions between human gut microbes, the host and invading pathogens during infection processes. Their research sought to identify and functionally characterize non-coding RNA molecules (ncRNA) and RNA-binding proteins in pathogens, the host and its microbiota to advance the development of targeted RNA-based diagnostics and therapeutics.
Their work focused on the transcriptome of the human gut commensal Bacteroides thetaiotaomicron in the context of host niche colonization in the human colon. They developed a sophisticated in vitro model of the human gut with cell organoids and utilizing RNA sequencing technologies, such as Dual RNA-seq and differential RNA-seq. The group's overarching goal was to find novel targets for the treatment and prevention of enteric infections based on RNA mechanisms.
Team members

Prof Alexander Westermann, Affiliated Scientist
Group Leader

Dr Ann-Sophie Rüttiger
Postdoc

Dr Lena Amend
Postdoc

Dr Takeshi Kanda
Postdoc

Dr Thomas Guest
Postdoc

Hoda Kooshapour
PhD Student

Nahyung Ko
PhD Student

Taís Franco de Carvalho
PhD Student

Lisa Pfeuffer
Technical Assistant
Research projects
Graphical Abstract
Alumni
Elise Bornet, PhD Student • Gohar Mädler, PhD Student • Gianluca Prezza, PhD Student • Sarah Reichardt, Technical Assistant
Publications
2025
Enabling next-generation anaerobic cultivation through biotechnology to advance functional microbiome research
Clavel T, Faber F, Groussin M, Haller D, Overmann J, Pauvert C, Poyet M, Selkrig J, Stecher B, Typas A, …, Wylensek D, Maier L (2025)
Nature Biotechnology 43 (6): 878-888
A Multicellular In Vitro Model of the Human Intestine with Immunocompetent Features Highlights Host-Pathogen Interactions During Early Salmonella Typhimurium Infection
Damigos S, Caliskan A, Wajant G, Giddins S, Moldovan A, Kuhn S, Putz E, Dandekar T, Rudel T, Westermann AJ, Zdzieblo D (2025)
Advanced Science 12 (9): e2411233
The global RNA-binding protein RbpB is a regulator of polysaccharide utilization in Bacteroides thetaiotaomicron
Rüttiger AS, Ryan D, Spiga L, Lamm-Schmidt V, Prezza G, Reichardt S, Langford M, Barquist L, Faber F, Zhu W, Westermann AJ (2025)
Nature Communications 16 (1): 208
Intramacrophage RIL-seq uncovers an RNA antagonist of the Salmonella virulence-associated small RNA PinT
Kooshapour H, Matera G, Venturini E, Metka L, Bischler T, Vogel J, Westermann AJ (2025)
Nucleic Acids Research 53 (22): gkaf1364
3'UTR-derived small RNA couples acid resistance to metabolic reprogramming of Salmonella within macrophages
Kanda T, Liu F, Kooshapour H, Reichardt S, Wang M, Icyishaka P, Obana N, Westermann AJ, Chao Y, Miyakoshi M (2025)
Nucleic Acids Research 53 (22): gkaf1371
Unveiling immune interference: how the dendritic cell response to co-infection with Aspergillus fumigatus is modulated by human cytomegalovirus and its virokine CMVIL-10
Heilig L, Bussemer L, Strobel L, Hünniger-Ast K, Kurzai O, Grothey A, Dölken L, Laib Sampaio K, Panagiotou G, Westermann AJ, …, Schäuble S, Löffler J (2025)
mBio 16 (11)
Low-input RNA-seq suggests metabolic specialization underlying morphological heterogeneity in a gut commensal bacterium
Bornet E, Prezza G, Cecchino L, Jenniches L, Behrends J, Tawk C, Huang KC, Strowig T, Vogel J, Barquist L, Saliba AE, Westermann AJ (2025)
Cell Reports 44 (6): 115844
A Small RNA Derived From the 5' End of the IS200 tnpA Transcript Regulates Multiple Virulence Regulons in Salmonella typhimurium
Trussler RS, Scherba NQ, Kooshapour H, Ellis MJ, Förstner KU, Albert M, Westermann AJ, Haniford DB (2025)
Molecular Microbiology 124 (5): 413-432
Functional characterization of the DUF1127-containing small protein YjiS of Salmonella Typhimurium
Venturini E, Maaß S, Bischler T, Becher D, Vogel J, Westermann AJ (2025)
microLife 6: uqae026
2024
Global analysis of the RNA-RNA interactome in Acinetobacter baumannii AB5075 uncovers a small regulatory RNA repressing the virulence-related outer membrane protein CarO
Hamrock FJ, Ryan D, Shaibah A, Ershova AS, Mogre A, Sulimani MM, Ben Taarit S, Reichardt S, Hokamp K, Westermann AJ, Kröger C (2024)
Nucleic Acids Research 52 (18): 11283–11300
Improved RNA stability estimation through Bayesian modeling reveals most Salmonella transcripts have subminute half-lives
Jenniches L, Michaux C, Popella L, Reichardt S, Vogel J, Westermann AJ, Barquist L (2024)
PNAS 121 (14): e2308814121
An expanded transcriptome atlas for Bacteroides thetaiotaomicron reveals a small RNA that modulates tetracycline sensitivity
Ryan D, Bornet E, Prezza G, Alampalli SV, Franco de Carvalho T, Felchle H, Ebbecke T, Hayward RJ, Deutschbauer AM, Barquist L, Westermann AJ (2024)
Nature Microbiology 9 (4): 1130-1144
CRISPR-based screening of small RNA modulators of bile susceptibility in Bacteroides thetaiotaomicron
Prezza G, Liao C, Reichardt S, Beisel CL, Westermann AJ (2024)
PNAS 121 (6): 1096
Non-professional efferocytosis of Salmonella-infected intestinal epithelial cells in the neonatal host
Zhang K, Repnik U, Diab N, Friske D, Pütz A, Bachmann AZ, Gubbi NMKP, Hensel M, Förstner KU, Westermann AJ, Dupont A, Hornef MW (2024)
The Journal of Experimental Medicine 221 (3)
Group B Streptococcus transcriptome when interacting with brain endothelial cells
Vollmuth N, Bridgers BE, Armstrong ML, Wood JF, Gildea AR, Espinal ER, Hooven TA, Barbieri G, Westermann AJ, Sauerwein T, …, Schubert-Unkmeir A, Kim BJ (2024)
Journal of Bacteriology 206 (6): e0008724
CRISPR Interference-Based Functional Small RNA Genomics
Prezza G, Westermann AJ (2024)
Methods in Molecular Biology 2741: 101-116
2023
Cytosolic Ptbp2 modulates axon growth in motoneurons through axonal localization and translation of Hnrnpr
Salehi S, Zare A, Prezza G, Bader J, Schneider C, Fischer U, Meissner F, Mann M, Briese M, Sendtner M (2023)
Nature Communications 14 (1): 4158
Systematically attenuating DNA targeting enables CRISPR-driven editing in bacteria
Collias D, Vialetto E, Yu J, Co K, Almási ÉDH, Rüttiger AS, Achmedov T, Strowig T, Beisel CL (2023)
Nature Communications 14 (1): 680
RNA recording in single bacterial cells using reprogrammed tracrRNAs
Jiao C, Reckstadt C, König F, Homberger C, Yu J, Vogel J, Westermann AJ, Sharma CM, Beisel CL (2023)
Nature Biotechnology 41 (8): 1107-1116
A primary cell-based in vitro model of the human small intestine reveals host olfactomedin 4 induction in response to Salmonella Typhimurium infection
Däullary T, Imdahl F, Dietrich O, Hepp L, Krammer T, Fey C, Neuhaus W, Metzger M, Vogel J, Westermann AJ, Saliba AE, Zdzieblo D (2023)
Gut Microbes 15 (1): 2186109
2022
TFEB induces mitochondrial itaconate synthesis to suppress bacterial growth in macrophages
Schuster EM, Epple MW, Glaser KM, Mihlan M, Lucht K, Zimmermann JA, Bremser A, Polyzou A, Obier N, Cabezas-Wallscheid N, …, Westermann AJ, Rambold AS (2022)
Nature Metabolism 4 (7): 856-866
The ambivalent role of Bacteroides in enteric infections
Bornet E, Westermann AJ (2022)
Trends in Microbiology 30 (2): 104-108
Comparative genomics provides structural and functional insights into Bacteroides RNA biology
Prezza G, Ryan D, Mädler G, Reichardt S, Barquist L, Westermann AJ (2022)
Molecular Microbiology 117 (1): 67-85
2021
SPI2 T3SS effectors facilitate enterocyte apical to basolateral transmigration of Salmonella-containing vacuoles in vivo
Fulde M, van Vorst K, Zhang K, Westermann AJ, Busche T, Huei YC, Welitschanski K, Froh I, Pägelow D, Plendl J, …, Repnik U, Hornef MW (2021)
Gut Microbes 13 (1): 1973836
Cross-species RNA-seq for deciphering host–microbe interactions
Westermann AJ, Vogel J (2021)
Nature Reviews Genetics 22 (6): 361–378
MAPS integrates regulation of actin-targeting effector SteC into the virulence control network of Salmonella small RNA PinT
Correia Santos S, Bischler T, Westermann AJ, Vogel J (2021)
Cell Reports 34 (5): 108722
2020
A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron
Ryan D, Jenniches L, Reichardt S, Barquist L, Westermann AJ (2020)
Nature Communications 11 (1): 3557
Dual RNA-seq of Orientia tsutsugamushi informs on host-pathogen interactions for this neglected intracellular human pathogen
Mika-Gospodorz B, Giengkam S, Westermann AJ, Wongsantichon J, Kion-Crosby W, Chuenklin S, Wang LC, Sunyakumthorn P, Sobota RM, Subbian S, …, Barquist L, Salje J (2020)
Nature Communications 11: 3363
An RNA-centric view on gut Bacteroidetes
Ryan D, Prezza G, Westermann AJ (2020)
Biological Chemistry 402 (1): 55-72
A global data-driven census of Salmonella small proteins and their potential functions in bacterial virulence
Venturini E, Svensson SL, Maaß S, Gelhausen R, Eggenhofer F, Li L, Cain AK, Parkhill J, Becher D, Backofen R, …, Westermann AJ, Vogel J (2020)
microLife 1 (1): 597
Triple RNA-Seq Reveals Synergy in a Human Virus-Fungus Co-infection Model
Seelbinder B, Wallstabe J, Marischen L, Weiss E, Wurster S, Page L, Löffler C, Bussemer L, Schmitt A, Wolf T, …, Schäuble S, Loeffler J (2020)
Cell Reports 33 (7): 108389
Amidochelocardin Overcomes Resistance Mechanisms Exerted on Tetracyclines and Natural Chelocardin
Hennessen F, Miethke M, Zaburannyi N, Loose M, Lukežic T, Bernecker S, Hüttel S, Jansen R, Schmiedel J, Fritzenwanker M, …, Herrmann J, Müller R (2020)
Antibiotics 9 (9): 619
Improved bacterial RNA-seq by Cas9-based depletion of ribosomal RNA reads
Prezza G, Heckel T, Dietrich S, Homberger C, Westermann AJ, Vogel J (2020)
RNA 26 (8): 1069-1078
An Advanced Human Intestinal Coculture Model Reveals Compartmentalized Host and Pathogen Strategies during Salmonella Infection
Schulte LN, Schweinlin M, Westermann AJ, Janga H, Santos SC, Appenzeller S, Walles H, Vogel J, Metzger M (2020)
mBio 11 (1): e03348-19
2019
Induced Pluripotent Stem Cell-Derived Brain Endothelial Cells as a Cellular Model to Study Neisseria meningitidis Infection
Martins Gomes SF, Westermann AJ, Sauerwein T, Hertlein T, Förstner KU, Ohlsen K, Metzger M, Shusta EV, Kim BJ, Appelt-Menzel A, Schubert-Unkmeir A (2019)
Frontiers in Microbiology 10: 1181
The Major RNA-Binding Protein ProQ Impacts Virulence Gene Expression in Salmonella enterica Serovar Typhimurium
Westermann AJ, Venturini E, Sellin ME, Förstner KU, Hardt W, Vogel J (2019)
mBio 10 (1): e02504-18
2018
Salmonella persisters undermine host immune defenses during antibiotic treatment
Stapels DA, Hill PW, Westermann AJ, Fisher RA, Thurston TL, Saliba AE, Blommestein I, Vogel J, Helaine S (2018)
Science 362 (6419): 1156-1160
Toward Cell Type-Specific In Vivo Dual RNA-Seq
Frönicke L, Bronner DN, Byndloss MX, McLaughlin B, Bäumler AJ, Westermann AJ (2018)
Methods In Enzymology 612: 505-522
Regulatory RNAs in Virulence and Host-Microbe Interactions
Westermann AJ (2018)
Microbiology Spectrum 6 (4): 305-337
MetaMap: an atlas of metatranscriptomic reads in human disease-related RNA-seq data
Simon LM, Karg S, Westermann AJ, Engel M, Elbehery AH, Hense B, Heinig M, Deng L, Theis FJ (2018)
GigaScience 7 (6): giy070
CRP-cAMP mediates silencing of Salmonella virulence at the post-transcriptional level
El Mouali Y, Gaviria-Cantin T, Sánchez-Romero MA, Gibert M, Westermann AJ, Vogel J, Balsalobre C (2018)
PLOS Genetics 14 (6): e1007401
Host-Pathogen Transcriptomics by Dual RNA-Seq
Westermann AJ, Vogel J (2018)
In: Arluison V., Valverde C. (eds) Bacterial Regulatory RNA, Methods in Molecular Biology 1737: 59-75
2017
P5SM suicide exon for regulating gene expression
Hammond MC, Westermann AJ, Qin Q (2017)
Patent (US9637750B2)
Einzelzell-RNA-Sequenzierung beleuchtet den Infektionsprozess
Saliba AE, Westermann AJ, Vogel J (2017)
BIOspektrum 23 (5): 525-528
RNA target profiles direct the discovery of virulence functions for the cold-shock proteins CspC and CspE
Michaux C, Holmqvist E, Vasicek E, Sharan M, Barquist L, Westermann AJ, Gunn JS, Vogel J (2017)
PNAS 114 (26): 6824-6829
Resolving host-pathogen interactions by dual RNA-seq
Westermann AJ, Barquist L, Vogel J (2017)
PLOS Pathogens 13 (2): e1006033
2016
Single-cell RNA-seq ties macrophage polarization to growth rate of intracellular Salmonella
Saliba AE, Li L, Westermann AJ, Appenzeller S, Stapels DA, Schulte LN, Helaine S, Vogel J (2016)
Nature Microbiology 2: 16206
Dual RNA-seq unveils noncoding RNA functions in host-pathogen interactions
Westermann AJ, Forstner KU, Amman F, Barquist L, Chao Y, Schulte LN, Muller L, Reinhardt R, Stadler PF, Vogel J (2016)
Nature 529 (7587): 496-501
An NK Cell Perforin Response Elicited via IL-18 Controls Mucosal Inflammation Kinetics during Salmonella Gut Infection
Müller AA, Dolowschiak T, Sellin ME, Felmy B, Verbree C, Gadient S, Westermann AJ, Vogel J, LeibundGut-Landmann S, Hardt W (2016)
PLOS Pathogens 12 (6): e1005723
Molecular phenotyping of infection-associated small non-coding RNAs
Barquist L, Westermann AJ, Vogel J (2016)
Philosophical Transactions of the Royal Society of London B: Biological Sciences 371 (1707): 20160081
2015
Dual 3'Seq using deepSuperSAGE uncovers transcriptomes of interacting Salmonella enterica Typhimurium and human host cells
Afonso-Grunz F, Hoffmeier K, Müller S, Westermann AJ, Rotter B, Vogel J, Winter P, Kahl G (2015)
BMC Genomics 16: 323
2014
Single-cell RNA-seq: advances and future challenges
Saliba AE, Westermann AJ, Gorski SA, Vogel J (2014)
Nucleic Acids Research 42 (14): 8845-60
2013
Differential activation and functional specialization of miR-146 and miR-155 in innate immune sensing
Schulte LN, Westermann AJ, Vogel J (2013)
Nucleic Acids Research 41 (1): 542-53
2012
Dual RNA-seq of pathogen and host
Westermann AJ, Gorski SA, Vogel J (2012)
Nature Reviews Microbiology 10 (9): 618-30
Transgene regulation in plants by alternative splicing of a suicide exon
Hickey SF, Sridhar M, Westermann AJ, Qin Q, Vijayendra P, Liou G, Hammond MC (2012)
Nucleic Acids Research 40 (10): 4701-10
2010
Polypyrimidine tract-binding protein homologues from Arabidopsis underlie regulatory circuits based on alternative splicing and downstream control
Stauffer E, Westermann A, Wagner G, Wachter A (2010)
The Plant journal : for cell and molecular biology 64 (2): 243-55