Prof Jörg Vogel
RNA Biology of Bacterial Infections
Our research and approach
The research group led by Jörg Vogel investigates the RNA world of microbes during infection and in the context of their natural habitat. Using high-resolution methods, they seek to understand when, how and why bacteria use RNA-based mechanisms to regulate their genes, and exploit this knowledge to develop RNA-based therapeutic approaches.
The use of broad-spectrum antibiotics causes collateral damage to the surrounding microbiota and is increasingly challenged by antimicrobial resistance. Precision medicine, such as programmable RNA antibiotics, offers the possibility to overcome such issues, but requires a comprehensive understanding of the major RNA-based pathways and mechanisms during infection.
Jörg Vogel's group explores the RNA biology, namely non-coding RNA and RNA-binding proteins, of bacterial pathogens and members of the human microbiome, as well as phage-host interactions. They develop deep sequencing-based techniques to capture the RNA world of microbes, ideally at the single-cell level, and seek to elucidate how bacteria use RNA as a regulator during infection.
The group's projects focus on several microorganisms, including Salmonella Typhimurium and Fusobacterium nucleatum, an anaerobic microbe associated with colorectal cancer. Their work combines cutting-edge throughput technologies, including single-cell RNA-seq, dual RNA-seq, or Grad-seq, with comprehensive molecular microbiology approaches. Their overarching goal is to exploit findings to advance the development of RNA-based therapies that target pathogens and manipulate the microbiota with precision.
Team members

Prof Jörg Vogel
Group Leader

Dr Gauthier Roy
Postdoc

Dr Manon Janet-Maitre
Postdoc

Dr Paramita Sarkar
Postdoc

Dr Toby Wilkinson
Postdoc

Dr Linda Popella
Postdoc (on parental leave)

Hanna Weidt
PhD Student

Julia Seidel
PhD Student

Leandro Buhlmann
PhD Student

Sarah Nentwich
PhD Student

Valentina Cosi
PhD Student

Victoria Gebler
PhD Student

Vincent Lau
PhD Student

Johanna Krzok
Technical Assistant

Luise Klos
Technical Assistant

Dr Esther Hauschild
Technical Assistant (on parental leave)

Dr Anke Sparmann
Scientific Writer
Research projects
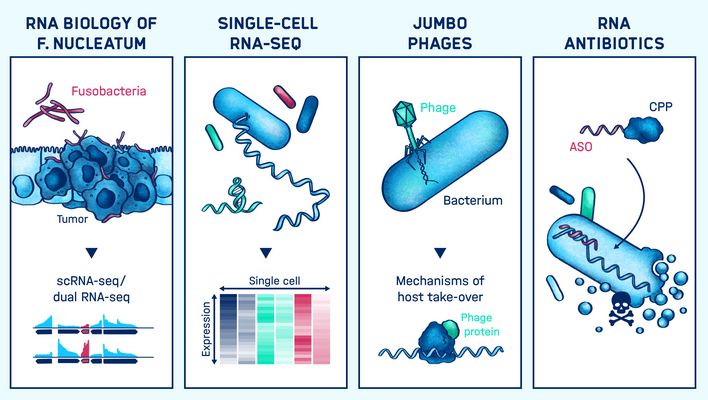
Graphical Abstract
Introducing the research group
Discovery of functional RNAs in the microbiome
The major classes of bacterial noncoding RNA are known only for a few species, meaning that the RNA biology of the vast majority of microbes that are important for human health remains unexplored. Exploiting RNA as a programmable antibiotic to treat bacterial infections and dysbiosis requires that we comprehensively understand the major RNA-based pathways and mechanisms in these species. Over the years, we have pioneered a number of RNA deep sequencing-based methods such as differential RNA-seq (Sharma et al. 2010 Nature) or Dual RNA-seq (Westermann et al. 2016 Nature) to understand which and when RNAs are expressed during the course of an infection. Moreover, with Grad-seq (Smirnov et al. 2016 PNAS) we can now classify potentially all cellular transcripts in an unbiased manner based on their biochemical behaviour. Using these approaches, we are beginning to chart the full cosmos of noncoding RNA functions in the human microbiome.

New RNA-binding proteins
Noncoding RNA molecules generally interact with RNA-binding proteins (RBPs) to become functional in the cellular environment. Decades of work in eukaryotes have established RBPs as a pillar of post-transcriptional gene control, with numbers ranging from the hundreds to thousands in vertebrates, invertebrates and yeast cells. Ironically, we know relatively little about the number of RBPs and the mechanisms by which they function in the organisms that seem the easiest to analyse: bacteria. Although bacteria extensively regulate gene expression at the RNA level, our knowledge of global RBPs remains limited to the small RNA chaperone Hfq and the translational repressor CsrA. However, increasing evidence, including the serendipitous discovery of CRISPR-Cas systems, suggests that many RBPs await discovery especially in less well studied microbes.
We have discovered the “osmoregulatory” protein ProQ as a new abundant RBP which binds hundreds cellular RNAs in Salmonella and E. coli (Smirnov et al. 2016 PNAS), including various small RNAs. Our results for ProQ suggest new molecular mechanisms of bacterial gene control at mRNA 3’ ends and a novel regulatory network whereby ProQ recognizes RNA structure rather than primary sequence (Smirnov et al. 2017 EMBO J, Holmqvist et al. 2018 Molecular Cell). We are also addressing the fundamental principles involved in RNA recognition by trying to understand how members of this RBP family use the FinO domain for either highly specific or global RNA recognition in the same cellular environment. Continuing with RBP discovery, we are now exploring the microbiome for new RBPs.

Single-cell RNA biology
The development of single-cell RNA-seq and its ability to describe the gene expression profiles of individual cells rather than the average cellular population is rapidly changing our view of many aspects of biology, including the discovery of new cell types and functions. Single-cell RNA-seq is also emerging as a powerful tool to determine cell-to-cell variability in bacterial pathogenesis, however, technical issues have restricted its use to the analysis of eukaryotic transcriptomes. For example, we have studied how heterogeneous intracellular growth of Salmonella drives the polarization state of individually infected macrophages (Saliba et al. 2016 Nature Microbiology).
Our grand challenge is to fully establish single-cell RNA-seq for bacteria, which is currently hampered by the small amounts of cellular RNA present and the lack of a polyA tail for easy cDNA priming as in eukaryotes. Transcriptomes from single bacteria will provide insight into the basis for phenotypic resistance to antibiotic treatment, stochastic persister formation, and phenotypic diversification for bet hedging, poorly understood phenomena that are threatening our ability to successfully treat bacterial infections. It will also be the next step for developing single-cell Dual RNA-seq to simultaneously study individual host-pathogen interactions. Going beyond simple single-cell transcriptomics, our long-term ambition is to develop robust methodologies for routine high-throughput RNA biochemistry to study post-transcriptional control in single bacterial cells.

The RNA route to manipulation of the microbiota
Our body is colonized by a vast array of bacteria, which together form the human microbiome. The gut alone harbours more than 1,000 bacterial species. An understanding of their individual or synergistic contributions to human health and disease demands a means to interfere and perturb their functions on the species level, with a selectivity that is not possible using currently available antibiotics.
Our ultimate goal is precision manipulation of the human microbiome using “programmable antibiotics” in the form of RNA-directed antisense oligonucleotides (ASOs). The same approach can be taken to target antibiotic-resistant pathogens as an alternative to standard antibiotics. These ASOs are coupled to small peptides that carry them inside the bacteria to silence the mRNAs of essential genes. There is already proof-of-principle for this exciting concept, but many open questions remain. What does it take to make these ASOs truly species-specific; how can we minimize potential off-target effects; what are the most effective peptides for delivery; what about the host response and bacterial resistance mechanisms? Since there is unlikely to be a one case-fits-all solution for all microbiome species, this project can only succeed by integrating diverse expertise from RNA biology, microbiology, immunology, single-cell biology, and pharmacy.

In focus
Oral microbe which speeds up growth of cancer cells
The oral cavity germ Fusobacterium nucleatum is known to speed up the growth of human carcinomas, for example in the intestine or breast. In a joint study, scientists from Vogel's labs at the Helmholtz Institute Würzburg and the Institute for Molecular Infection Biology have mapped the RNA molecules of five clinically relevant strains of this adaptable bacterium. The findings could help develop new therapies for various cancers and were published in the journal Nature Microbiology.
Alumni
Irene Calvin, Technical Assistant • Kotaro Chihara, Postdoc • Svetlana Durica-Mitic, Postdoc • Milan Gerovac, Postdoc • Chandradhish Ghosh, Postdoc • Christina Homberger, PhD Student • Jakob Jung, PhD Student & Postdoc • Tobias Kerrinnes, Lab Coordinator • Gianluca Matera, PhD Student • Petia Petkova-Datchary, Postdoc • Sushila Pisano, PhD Student • Barbara Plaschke, Technical Assistant • Falk Ponath, Postdoc • Phuong Thao Do, Technical Assistant • Laura Vogel, Technical Assistant • Yan Zhu, PhD Student & Postdoc • Anna Zweyer, Technical Assistant
Publications
2026
Gleich und doch verschieden: bakterieller Genexpression auf der Spur
Imdahl F, Homberger C, Vogel J (2026)
BIOspektrum 32 (3): 263-265
Translation-dependent degradation of cas12 mRNA triggered by an anti-CRISPR
Marino ND, Talaie A, Gerovac M, Rodriguez JL, Schmidt AD, Astmann TJ, Carion H, Taylor AF, Liliedahl J, Haniyur S, …, Vogel J, Bondy-Denomy J (2026)
Nature (Online ahead of print)
A 2025 meeting report from the Venice lagoon: International graduate program "RNAmed - Future Leaders in RNA-based Medicine" meets on San Servolo
Vogel J, Kamm C, Gerolimetto G, Meel P, Trubert A, Rademacker S, Fröschel C, Vogel J, Kamm C, Gerolimetto G, …, Rademacker S, Fröschel C (2026)
RNA (Online ahead of print)
Developing antisense oligomer biotics "asobiotics" as precision antibacterials: designs, strategies, and considerations for future success
Moustafa DA, Ghosh C, Jung J, Nanayakkara A, Sapkota M, Nagamatsu K, Geller B, Vogel J, Goldberg JB, Greenberg DE (2026)
FEMS Microbiology Reviews 50: fuaf059
2025
Pan-modification profiling facilitates a cross-evolutionary dissection of the thermoregulated ribosomal epitranscriptome
Garcia-Campos MA, Georgeson J, Nir R, Reichelt R, Fluke KA, Matzov D, Iyer V, Burkhart BW, Lui L, Kustanovich A, …, Shalev-Benami M, Schwartz S (2025)
Cell 188 (24): 6825-6844.e28
Programmable antisense oligomers for phage functional genomics
Gerovac M, Buhlmann L, Zhu Y, Ðurica-Mitic S, Rech V, Carien S, Gräfenhan T, Popella L, Vogel J, Gerovac M, …, Popella L, Vogel J (2025)
Nature 646 (8087): 1195-1203
High-Throughput Tiling of Essential mRNAs Increases Potency of Antisense Antibiotics
Danti G, Popella L, Vogel J, Maric HM (2025)
Advanced Science 12 (28): e2504284
Intramacrophage RIL-seq uncovers an RNA antagonist of the Salmonella virulence-associated small RNA PinT
Kooshapour H, Matera G, Venturini E, Metka L, Bischler T, Vogel J, Westermann AJ (2025)
Nucleic Acids Research 53 (22): gkaf1364
Fusobacterium nucleatum interacts with cancer-associated fibroblasts to promote colorectal cancer
Karta J, Meyers M, Rodriguez F, Koncina E, Gilson C, Klein E, Gabola M, Benzarti M, Pérez Escriva P, Molina Tijeras JA, …, Wilmes P, Letellier E (2025)
The EMBO Journal 44 (19)
Low-input RNA-seq suggests metabolic specialization underlying morphological heterogeneity in a gut commensal bacterium
Bornet E, Prezza G, Cecchino L, Jenniches L, Behrends J, Tawk C, Huang KC, Strowig T, Vogel J, Barquist L, Saliba AE, Westermann AJ (2025)
Cell Reports 44 (6): 115844
A Small RNA Derived From the 5' End of the IS200 tnpA Transcript Regulates Multiple Virulence Regulons in Salmonella typhimurium
Trussler RS, Scherba NQ, Kooshapour H, Ellis MJ, Förstner KU, Albert M, Westermann AJ, Haniford DB (2025)
Molecular Microbiology 124 (5): 413-432
An antisense oligomer conjugate with unpredicted bactericidal activity against Fusobacterium nucleatum
Cosi V, Jung J, Popella L, Ponath F, Ghosh C, Barquist L, Vogel J (2025)
mBio 16 (6): e0052425
RNA toehold switch-based reporter assay to assess bacterial uptake of antisense oligomers
Sarkar P, Popella L, Pérez-Jiménez S, Vogel J (2025)
mBio 16 (4): e0398324
A ribosome-interacting jumbophage protein associates with the phage nucleus to facilitate efficient propagation
Wannasrichan W, Krobthong S, Morgan CJ, Armbruster EG, Gerovac M, Yingchutrakul Y, Wongtrakoongate P, Vogel J, Aonbangkhen C, Nonejuie P, Pogliano J, Chaikeeratisak V (2025)
PLOS Pathogens 21 (2): e1012936
Functional characterization of the DUF1127-containing small protein YjiS of Salmonella Typhimurium
Venturini E, Maaß S, Bischler T, Becher D, Vogel J, Westermann AJ (2025)
microLife 6: uqae026
2024
The RNA landscape of the human commensal Segatella copri reveals a small RNA essential for gut colonization
El Mouali Y, Tawk C, Huang KD, Amend L, Lesker TR, Ponath F, Vogel J, Strowig T (2024)
Cell Host & Microbe 32 (11): 1910-1926.e6
Cooperation of regulatory RNA and the RNA degradosome in transcript surveillance
Bandyra KJ, Fröhlich KS, Vogel J, Rodnina M, Goyal A, Luisi BF (2024)
Nucleic Acids Research 52 (15): 9161-9173
Phage proteins target and co-opt host ribosomes immediately upon infection
Gerovac M, Chihara K, Wicke L, Böttcher B, Lavigne R, Vogel J (2024)
Nature Microbiology 9 (3): 787-800
Improved RNA stability estimation through Bayesian modeling reveals most Salmonella transcripts have subminute half-lives
Jenniches L, Michaux C, Popella L, Reichardt S, Vogel J, Westermann AJ, Barquist L (2024)
PNAS 121 (14): e2308814121
A global survey of small RNA interactors identifies KhpA and KhpB as major RNA-binding proteins in Fusobacterium nucleatum
Zhu Y, Ponath F, Cosi V, Vogel J (2024)
Nucleic Acids Research 52 (7): 3950-3970
Global Hfq-mediated RNA interactome of nitrogen starved Escherichia coli uncovers a conserved post-transcriptional regulatory axis required for optimal growth recovery
McQuail J, Matera G, Gräfenhan T, Bischler T, Haberkant P, Stein F, Vogel J, Wigneshweraraj S (2024)
Nucleic Acids Research 52 (5): 2323-2339
Refining the transcriptional landscapes for distinct clades of virulent phages infecting Pseudomonas aeruginosa
Putzeys L, Wicke L, Boon M, van Noort V, Vogel J, Lavigne R (2024)
microLife 5: uqae002
Transcriptome fine-mapping in Fusobacterium nucleatum reveals FoxJ, a new σE-dependent small RNA with unusual mRNA activation activity
Ponath F, Zhu Y, Vogel J (2024)
mBio 15 (4): e0353623
A comparative analysis of peptide-delivered antisense antibiotics employing diverse nucleotide mimics
Ghosh C, Popella L, Dhamodharan V, Jung J, Dietzsch J, Barquist L, Höbartner C, Vogel J (2024)
RNA 30 (6): 624-643
Exploring the transcriptional landscape of phage-host interactions using novel high-throughput approaches
Putzeys L, Wicke L, Brandão A, Boon M, Pires DP, Azeredo J, Vogel J, Lavigne R, Gerovac M (2024)
Current Opinion in Microbiology 77: 102419
2023
Brain-to-gut trafficking of alpha-synuclein by CD11c+ cells in a mouse model of Parkinson's disease
McFleder RL, Makhotkina A, Groh J, Keber U, Imdahl F, Peña Mosca J, Peteranderl A, Wu J, Tabuchi S, Hoffmann J, …, Volkmann J, Ip CW (2023)
Nature Communications 14 (1): 7529
A SAM analogue-utilizing ribozyme for site-specific RNA alkylation in living cells
Okuda T, Lenz AK, Seitz F, Vogel J, Höbartner C (2023)
Nature Chemistry 15 (11): 1523-1531
RNA-based medicine: from molecular mechanisms to therapy
Sparmann A, Vogel J (2023)
The EMBO Journal 42 (21): e114760
Th17.1 cell driven sarcoidosis-like inflammation after anti-BCMA CAR T cells in multiple myeloma
Leipold AM, Werner RA, Düll J, Jung P, John M, Stanojkovska E, Zhou X, Hornburger H, Ruckdeschel A, Dietrich O, …, Saliba AE, Rasche L (2023)
Leukemia 37 (3): 650-658
RNA recording in single bacterial cells using reprogrammed tracrRNAs
Jiao C, Reckstadt C, König F, Homberger C, Yu J, Vogel J, Westermann AJ, Sharma CM, Beisel CL (2023)
Nature Biotechnology 41 (8): 1107-1116
EAM highlights in FEMS 2023: from the Petri dish to planet Earth
Vogel J, de Lorenzo V (2023)
microLife 4: uqad045
Seropositivity of Borrelia burgdorferi s.l. in Germany-an analysis across four German National Cohort (NAKO) study sites
Hassenstein MJ, Pischon T, Karch A, Peters A, Kerrinnes T, Teismann H, Schneider A, Thierry S, Moreno Velásquez I, Janke J, Kemmling Y, Castell S (2023)
Scientific Reports 13 (1): 21087
Grad-seq analysis of Enterococcus faecalis and Enterococcus faecium provides a global view of RNA and protein complexes in these two opportunistic pathogens
Michaux C, Gerovac M, Hansen EE, Barquist L, Vogel J (2023)
microLife 4: uqac027
Gut Microbiota Contribution to Weight-Independent Glycemic Improvements after Gastric Bypass Surgery
Hankir MK, Kovatcheva-Datchary P, Springer R, Hoffmann A, Vogel J, Seyfried F, Arora T (2023)
Microbiology Spectrum 11 (3): e0510922
Improved Bacterial Single-Cell RNA-Seq through Automated MATQ-Seq and Cas9-Based Removal of rRNA Reads
Homberger C, Hayward RJ, Barquist L, Vogel J (2023)
mBio 14 (2): e0355722
A primary cell-based in vitro model of the human small intestine reveals host olfactomedin 4 induction in response to Salmonella Typhimurium infection
Däullary T, Imdahl F, Dietrich O, Hepp L, Krammer T, Fey C, Neuhaus W, Metzger M, Vogel J, Westermann AJ, Saliba AE, Zdzieblo D (2023)
Gut Microbes 15 (1): 2186109
Seroprevalence of Hepatitis E virus in children and adolescents living in urban Bogotá: An explorative cross-sectional study
Fernández Villalobos NV, Kessel B, Torres Páez JC, Strömpl J, Kerrinnes T, de la Hoz Restrepo FP, Strengert M, Krause G (2023)
Frontiers in Public Health 11: 981172
Design and off-target prediction for antisense oligomers targeting bacterial mRNAs with the MASON webserver
Jung JJ, Popella L, Do PT, Pfau P, Vogel J, Barquist L (2023)
RNA 299 (5): 570-583
A MATQ-seq-Based Protocol for Single-Cell RNA-seq in Bacteria
Homberger C, Saliba AE, Vogel J (2023)
Methods in Molecular Biology 2584: 105-121
2022
Hypothalamic integrity is necessary for sustained weight loss after bariatric surgery: A prospective, cross-sectional study
Dischinger U, Kötzner L, Kovatcheva-Datchary P, Kleinschmidt H, Haas C, Perez J, Presek C, Koschker AC, Miras AD, Hankir MK, …, Herrmann MJ, Seyfried F (2022)
Metabolism: Clinical and Experimental 138: 155341
An unusual mode of baseline translation adjusts cellular protein synthesis capacity to metabolic needs
Schneider C, Erhard F, Binotti B, Buchberger A, Vogel J, Fischer U (2022)
Cell Reports 41 (2): 111467
INRI-seq enables global cell-free analysis of translation initiation and off-target effects of antisense inhibitors
Hör J, Jung J, Ðurica-Mitic S, Barquist L, Vogel J (2022)
Nucleic Acids Research 50 (22): e128
Expanding the genetic toolkit helps dissect a global stress response in the early-branching species Fusobacterium nucleatum
Ponath F, Zhu Y, Cosi V, Vogel J (2022)
PNAS 119 (40): e2201460119
TFEB induces mitochondrial itaconate synthesis to suppress bacterial growth in macrophages
Schuster EM, Epple MW, Glaser KM, Mihlan M, Lucht K, Zimmermann JA, Bremser A, Polyzou A, Obier N, Cabezas-Wallscheid N, …, Westermann AJ, Rambold AS (2022)
Nature Metabolism 4 (7): 856-866
Comprehensive analysis of PNA-based antisense antibiotics targeting various essential genes in uropathogenic Escherichia coli
Popella L, Jung J, Do PT, Hayward RJ, Barquist L, Vogel J (2022)
Nucleic Acids Research 50 (11): 6435–6452
An overview of gene regulation in bacteria by small RNAs derived from mRNA 3' ends
Ponath F, Hör J, Vogel J (2022)
FEMS Microbiology Reviews 46 (5): fuac017
Global RNA interactome of Salmonella discovers a 5' UTR sponge for the MicF small RNA that connects membrane permeability to transport capacity
Matera G, Altuvia Y, Gerovac M, El Mouali Y, Margalit H, Vogel J (2022)
Molecular Cell 82 (3): 629-644.e4
The DNA polymerase of bacteriophage YerA41 replicates its T-modified DNA in a primer-independent manner
Gomez-Raya-Vilanova MV, Leskinen K, Bhattacharjee A, Virta P, Rosenqvist P, Smith JLR, Bayfield OW, Homberger C, Kerrinnes T, Vogel J, Pajunen MI, Skurnik M (2022)
Nucleic Acids Research 50 (7): 3985-3997
Ushering in a new era of single-cell transcriptomics in bacteria
Homberger C, Barquist L, Vogel J (2022)
microLife 3: uqac020
Global profiling of the RNA and protein complexes of Escherichia coli by size exclusion chromatography followed by RNA sequencing and mass spectrometry (SEC-seq)
Chihara K, Gerovac M, Hör J, Vogel J (2022)
RNA 29 (1): 123-39
Seroepidemiology of Borrelia burgdorferi s.l. among German National Cohort (NAKO) Participants, Hanover
Hassenstein MJ, Janzen I, Krause G, Harries M, Melhorn V, Kerrinnes T, Kemmling Y, Castell S (2022)
Microorganisms 10 (11): 2286
Regional seropositivity for Borrelia burgdorferi and associated risk factors: findings from the Rhineland Study, Germany
Coors A, Hassenstein MJ, Krause G, Kerrinnes T, Harries M, Breteler MMB, Castell S (2022)
Parasites & Vectors 15 (1): 241
Cellular RNA Targets of Cold Shock Proteins CspC and CspE and Their Importance for Serum Resistance in Septicemic Escherichia coli
Yair Y, Michaux C, Biran D, Bernhard J, Vogel J, Barquist L, Ron EZ (2022)
mSystems 7 (4): e0008622
Comparative Magnitude and Persistence of Humoral SARS-CoV-2 Vaccination Responses in the Adult Population in Germany
Dulovic A, Kessel B, Harries M, Becker M, Ortmann J, Griesbaum J, Jüngling J, Junker D, Hernandez P, Gornyk D, …, Schneiderhan-Marra N, Strengert M (2022)
Frontiers in Immunology 13: 828053
Innovative developments and emerging technologies in RNA therapeutics
Halloy F, Biscans A, Bujold KE, Debacker A, Hill AC, Lacroix A, Luige O, Strömberg R, Sundstrom L, Vogel J, Ghidini A (2022)
RNA Biology 19 (1): 313-332
2021
SPI2 T3SS effectors facilitate enterocyte apical to basolateral transmigration of Salmonella-containing vacuoles in vivo
Fulde M, van Vorst K, Zhang K, Westermann AJ, Busche T, Huei YC, Welitschanski K, Froh I, Pägelow D, Plendl J, …, Repnik U, Hornef MW (2021)
Gut Microbes 13 (1): 1973836
RNA landscape of the emerging cancer-associated microbe Fusobacterium nucleatum
Ponath F, Tawk C, Zhu Y, Barquist L, Faber F, Vogel J (2021)
Nature Microbiology 6 (8): 1007-1020
In vivo targets of Salmonella FinO include a FinP-like small RNA controlling copy number of a cohabitating plasmid
El Mouali Y, Gerovac M, Mineikaite R, Vogel J (2021)
Nucleic Acids Research 49 (9): 5319-5335
Analysis of the RNA and Protein Complexome by Grad-seq
Hör J, Vogel J (2021)
Methods in Molecular Biology 2300: 183-201
Global RNA profiles show target selectivity and physiological effects of peptide-delivered antisense antibiotics
Popella L, Jung J, Popova K, Ðurica-Mitic S, Barquist L, Vogel J (2021)
Nucleic Acids Research 49 (8): 4705-4724
Cross-species RNA-seq for deciphering host–microbe interactions
Westermann AJ, Vogel J (2021)
Nature Reviews Genetics 22 (6): 361–378
Opposing Wnt signals regulate cervical squamocolumnar homeostasis and emergence of metaplasia
Chumduri C, Gurumurthy RK, Berger H, Dietrich O, Kumar N, Koster S, Brinkmann V, Hoffmann K, Drabkina M, Arampatzi P, …, Saliba AE, Meyer TF (2021)
Nature Cell Biology 23 (2): 184–197
Grad-seq identifies KhpB as a global RNA-binding protein in Clostridioides difficile that regulates toxin production
Lamm-Schmidt V, Fuchs M, Sulzer J, Gerovac M, Hör J, Dersch P, Vogel J, Faber F (2021)
microLife 2: uqab004
Impact of pseudouridylation, substrate fold, and degradosome organization on the endonuclease activity of RNase E
Islam MS, Bandyra KJ, Chao Y, Vogel J, Luisi BF (2021)
RNA 27 (11): 1339-1352
Scanning mutagenesis of RNA-binding protein ProQ reveals a quality control role for the Lon protease
El Mouali Y, Ponath F, Scharrer V, Wenner N, Hinton JCD, Vogel J (2021)
RNA 27 (12): 1512-1527
SARS-CoV-2 Seroprevalence in Germany
Gornyk D, Harries M, Glöckner S, Strengert M, Kerrinnes T, Heise JK, Maaß H, Ortmann J, Kessel B, Kemmling Y, Lange B, Krause G (2021)
Deutsches Ärzteblatt international 118 (48): 824-831
An RNA-centric global view of Clostridioides difficile reveals broad activity of Hfq in a clinically important gram-positive bacterium
Fuchs M, Lamm-Schmidt V, Sulzer J, Ponath F, Jenniches L, Kirk JA, Fagan RP, Barquist L, Vogel J, Faber F (2021)
PNAS 118 (25): e2103579118
The World of Stable Ribonucleoproteins and Its Mapping With Grad-Seq and Related Approaches
Gerovac M, Vogel J, Smirnov A (2021)
Frontiers in Molecular Biosciences 8: 661448
A Grad-seq View of RNA and Protein Complexes in Pseudomonas aeruginosa under Standard and Bacteriophage Predation Conditions
Gerovac M, Wicke L, Chihara K, Schneider C, Lavigne R, Vogel J (2021)
mBio 12 (1): e03454-20
MAPS integrates regulation of actin-targeting effector SteC into the virulence control network of Salmonella small RNA PinT
Correia Santos S, Bischler T, Westermann AJ, Vogel J (2021)
Cell Reports 34 (5): 108722
A genome-wide transcriptomic analysis of embryos fathered by obese males in a murine model of diet-induced obesity
Bernhardt L, Dittrich M, El-Merahbi R, Saliba AE, Müller T, Sumara G, Vogel J, Nichols-Burns S, Mitchell M, Haaf T, El Hajj N (2021)
Scientific Reports 11: 1979
Introducing differential RNA-seq mapping to track the early infection phase for Pseudomonas phage ɸKZ
Wicke L, Ponath F, Coppens L, Gerovac M, Lavigne R, Vogel J (2021)
RNA Biology 18 (8): 1099-1110
2020
Longitudinal Multi-omics Analyses Identify Responses of Megakaryocytes, Erythroid Cells, and Plasmablasts as Hallmarks of Severe COVID-19
Bernardes JP, Mishra N, Tran F, Bahmer T, Best L, Blase JI, Bordoni D, Franzenburg J, Geisen U, Josephs-Spaulding J, …, Schultze JL, Rosenstiel P (2020)
Immunity 53 (6): 1296-1314.e9
Grad-seq shines light on unrecognized RNA and protein complexes in the model bacterium Escherichia coli
Hör J, Di Giorgio S, Gerovac M, Venturini E, Förstner KU, Vogel J (2020)
Nucleic Acids Research 48 (16): 9301-9319
LifeTime and improving European healthcare through cell-based interceptive medicine
Rajewsky N, Almouzni G, Gorski SA, Aerts S, Amit I, Bertero MG, Bock C, Bredenoord AL, Cavalli G, Chiocca S, …, Vidal M, Voet T (2020)
Nature 587 (7834): 377-386
Single-cell RNA-sequencing reports growth-condition-specific global transcriptomes of individual bacteria
Imdahl F, Vafadarnejad E, Homberger C, Saliba AE, Vogel J (2020)
Nature Microbiology 5 (10): 1202–1206
Severe COVID-19 is marked by a dysregulated myeloid cell compartment
Schulte-Schrepping J, Reusch N, Paclik D, Baßler K, Schlickeiser S, Zhang B, Krämer B, Krammer T, Brumhard S, Bonaguro L, …, Saliba AE, Sander LE (2020)
Cell 182 (6): 1419-1440
Dual RNA-seq of Orientia tsutsugamushi informs on host-pathogen interactions for this neglected intracellular human pathogen
Mika-Gospodorz B, Giengkam S, Westermann AJ, Wongsantichon J, Kion-Crosby W, Chuenklin S, Wang LC, Sunyakumthorn P, Sobota RM, Subbian S, …, Barquist L, Salje J (2020)
Nature Communications 11: 3363
Breast cancer colonization by Fusobacterium nucleatum accelerates tumor growth and metastatic progression
Parhi L, Alon-Maimon T, Sol A, Nejman D, Shhadeh A, Fainsod-Levi T, Yajuk O, Isaacson B, Abed J, Maalouf N, …, Straussman R, Bachrach G (2020)
Nature Communications 11: 3259
The minimal meningococcal ProQ protein has an intrinsic capacity for structure-based global RNA recognition
Bauriedl S, Gerovac M, Heidrich N, Bischler T, Barquist L, Vogel J, Schoen C (2020)
Nature Communications 11: 2823
Grad-seq in a Gram-positive bacterium reveals exonucleolytic sRNA activation in competence control
Hör J, Garriss G, Di Giorgio S, Hack LM, Vanselow JT, Förstner KU, Schlosser A, Henriques-Normark B, Vogel J (2020)
The EMBO Journal 39 (9): e103852
The conserved 3' UTR-derived small RNA NarS mediates mRNA crossregulation during nitrate respiration
Wang C, Chao Y, Matera G, Gao Q, Vogel J (2020)
Nucleic Acids Research 48 (4): 2126-2143
An RNA biology perspective on species-specific programmable RNA antibiotics
Vogel J (2020)
Molecular Microbiology 113 (3): 550-559
A global data-driven census of Salmonella small proteins and their potential functions in bacterial virulence
Venturini E, Svensson SL, Maaß S, Gelhausen R, Eggenhofer F, Li L, Cain AK, Parkhill J, Becher D, Backofen R, …, Westermann AJ, Vogel J (2020)
microLife 1 (1): 597
Single-Nucleotide RNA Maps for the Two Major Nosocomial Pathogens Enterococcus faecalis and Enterococcus faecium
Michaux C, Hansen EE, Jenniches L, Gerovac M, Barquist L, Vogel J (2020)
Frontiers in Cellular and Infection Microbiology 10: 600325
The SARS-CoV-2 RNA-protein interactome in infected human cells
Schmidt N, Lareau CA, Keshishian H, Ganskih S, Schneider C, Hennig T, Melanson R, Werner S, Wei Y, Zimmer M, …, Bodem J, Munschauer M (2020)
Nature Microbiology 6 (3): 339-353
Triple RNA-Seq Reveals Synergy in a Human Virus-Fungus Co-infection Model
Seelbinder B, Wallstabe J, Marischen L, Weiss E, Wurster S, Page L, Löffler C, Bussemer L, Schmitt A, Wolf T, …, Schäuble S, Loeffler J (2020)
Cell Reports 33 (7): 108389
Global discovery of bacterial RNA-binding proteins by RNase-sensitive gradient profiles reports a new FinO domain protein
Gerovac M, El Mouali Y, Kuper J, Kisker C, Barquist L, Vogel J (2020)
RNA 26 (10): 1448-1463
Amidochelocardin Overcomes Resistance Mechanisms Exerted on Tetracyclines and Natural Chelocardin
Hennessen F, Miethke M, Zaburannyi N, Loose M, Lukežic T, Bernecker S, Hüttel S, Jansen R, Schmiedel J, Fritzenwanker M, …, Herrmann J, Müller R (2020)
Antibiotics 9 (9): 619
Seropositivity for pathogens associated with chronic infections is a risk factor for all-cause mortality in the elderly: findings from the Memory and Morbidity in Augsburg Elderly (MEMO) Study
Zeeb M, Kerrinnes T, Cicin-Sain L, Guzman CA, Puppe W, Schulz TF, Peters A, Berger K, Castell S, Karch A (2020)
Geroscience 42: 1365-1376
Increasing storage stability of freeze-dried plasma using trehalose
Brogna R, Oldenhof H, Sieme H, Figueiredo C, Kerrinnes T, Wolkers WF (2020)
PLOS One 15 (6): e0234502
Improved bacterial RNA-seq by Cas9-based depletion of ribosomal RNA reads
Prezza G, Heckel T, Dietrich S, Homberger C, Westermann AJ, Vogel J (2020)
RNA 26 (8): 1069-1078
Trans-Acting Small RNAs and Their Effects on Gene Expression in Escherichia coli and Salmonella enterica
Hör J, Matera G, Vogel J, Gottesman S, Storz G (2020)
EcoSal Plus 9 (1): 10.1128/ecosalplus.ESP-0030-2019
Tracheal brush cells release acetylcholine in response to bitter tastants for paracrine and autocrine signaling
Hollenhorst MI, Jurastow I, Nandigama R, Appenzeller S, Li L, Vogel J, Wiederhold S, Althaus M, Empting M, Altmüller J, …, Saliba AE, Krasteva-Christ G (2020)
The FASEB Journal 34 (1): 316-332
An Advanced Human Intestinal Coculture Model Reveals Compartmentalized Host and Pathogen Strategies during Salmonella Infection
Schulte LN, Schweinlin M, Westermann AJ, Janga H, Santos SC, Appenzeller S, Walles H, Vogel J, Metzger M (2020)
mBio 11 (1): e03348-19
2019
An RNA Surprise in Bacterial Effector Mechanisms
Gerovac M, Vogel J (2019)
Cell Host & Microbe 26 (6): 709-711
Functional expansion of a TCA cycle operon mRNA by a 3' end-derived small RNA
Miyakoshi M, Matera G, Maki K, Sone Y, Vogel J (2019)
Nucleic Acids Research 47 (4): 2075-2088
Conditional Hfq Association with Small Noncoding RNAs in Pseudomonas aeruginosa Revealed through Comparative UV Cross-Linking Immunoprecipitation Followed by High-Throughput Sequencing
Chihara K, Bischler T, Barquist L, Monzon VA, Noda N, Vogel J, Tsuneda S (2019)
mSystems 4 (6): e00590-19
The CRISPR/Cas system in Neisseria meningitidis affects bacterial adhesion to human nasopharyngeal epithelial cells
Heidrich N, Hagmann A, Bauriedl S, Vogel J, Schoen C (2019)
RNA Biology 16 (4): 390-396
The Major RNA-Binding Protein ProQ Impacts Virulence Gene Expression in Salmonella enterica Serovar Typhimurium
Westermann AJ, Venturini E, Sellin ME, Förstner KU, Hardt W, Vogel J (2019)
mBio 10 (1): e02504-18
2018
Salmonella persisters undermine host immune defenses during antibiotic treatment
Stapels DA, Hill PW, Westermann AJ, Fisher RA, Thurston TL, Saliba AE, Blommestein I, Vogel J, Helaine S (2018)
Science 362 (6419): 1156-1160
Genome organization and DNA accessibility control antigenic variation in trypanosomes
Müller LS, Cosentino RO, Förstner KU, Guizetti J, Wedel C, Kaplan N, Janzen CJ, Arampatzi P, Vogel J, Steinbiss S, …, Sebra RP, Siegel TN (2018)
Nature 563 (7729): 121-125
Stress-induced host membrane remodeling protects from infection by non-motile bacterial pathogens
Tawk C, Nigro G, Rodrigues Lopes I, Aguilar C, Lisowski C, Mano M, Sansonetti P, Vogel J, Eulalio A (2018)
The EMBO Journal 37 (23): pii: e98529
RNA-binding proteins in bacteria
Holmqvist E, Vogel J (2018)
Nature Reviews Microbiology 16 (10): 601-615
Bacterial RNA Biology on a Genome Scale
Hör J, Gorski SA, Vogel J (2018)
Molecular Cell 70 (5): 785-799
Nuclear lncRNA stabilization in the host response to bacterial infection
Munschauer M, Vogel J (2018)
The EMBO Journal 37 (13): e99875
Global Maps of ProQ Binding In Vivo Reveal Target Recognition via RNA Structure and Stability Control at mRNA 3' Ends
Holmqvist E, Li L, Bischler T, Barquist L, Vogel J (2018)
Molecular Cell 70 (5): 971-982.e6
ANNOgesic: a Swiss army knife for the RNA-seq based annotation of bacterial/archaeal genomes
Yu S, Vogel J, Förstner KU (2018)
GigaScience 7 (9): giy096
CRP-cAMP mediates silencing of Salmonella virulence at the post-transcriptional level
El Mouali Y, Gaviria-Cantin T, Sánchez-Romero MA, Gibert M, Westermann AJ, Vogel J, Balsalobre C (2018)
PLOS Genetics 14 (6): e1007401
Host-Pathogen Transcriptomics by Dual RNA-Seq
Westermann AJ, Vogel J (2018)
In: Arluison V., Valverde C. (eds) Bacterial Regulatory RNA, Methods in Molecular Biology 1737: 59-75
2017
RNA-based recognition and targeting: sowing the seeds of specificity
Gorski SA, Vogel J, Doudna JA (2017)
Nature Reviews Molecular Cell Biology 18 (4): 215-228
APRICOT: an integrated computational pipeline for the sequence-based identification and characterization of RNA-binding proteins
Sharan M, Förstner KU, Eulalio A, Vogel J (2017)
Nucleic Acids Research 45 (11): e96
In Vivo Cleavage Map Illuminates the Central Role of RNase E in Coding and Non-coding RNA Pathways
Chao Y, Li L, Girodat D, Förstner KU, Said N, Corcoran C, Smiga M, Papenfort K, Reinhardt R, Wieden H, Luisi BF, Vogel J (2017)
Molecular Cell 65 (1): 39-51
Molecular mechanism of mRNA repression in trans by a ProQ‐dependent small RNA
Smirnov A, Wang C, Drewry LL, Vogel J (2017)
The EMBO Journal 36 (8): 1029-45
The primary transcriptome of Neisseria meningitidis and its interaction with the RNA chaperone Hfq
Heidrich N, Bauriedl S, Barquist L, Li L, Schoen C, Vogel J (2017)
Nucleic Acids Research 45 (10): 6147-6167
A systematic analysis of the RNA-targeting potential of secreted bacterial effector proteins
Tawk C, Sharan M, Eulalio A, Vogel J (2017)
Scientific Reports 7 (1): 9328
Einzelzell-RNA-Sequenzierung beleuchtet den Infektionsprozess
Saliba AE, Westermann AJ, Vogel J (2017)
BIOspektrum 23 (5): 525-528
Structure of the Escherichia coli ProQ RNA-binding protein
Gonzalez GM, Hardwick SW, Maslen SL, Skehel JM, Holmqvist E, Vogel J, Bateman A, Luisi BF, Broadhurst RW (2017)
RNA 23 (5): 696-711
New RNA-seq approaches for the study of bacterial pathogens
Saliba AE, Santos SC, Vogel J (2017)
Current Opinion in Microbiology 35: 78-87
Discovery of new RNA classes and global RNA-binding proteins
Smirnov A, Schneider C, Hör J, Vogel J (2017)
Current Opinion in Microbiology 39: 152-160
RNA target profiles direct the discovery of virulence functions for the cold-shock proteins CspC and CspE
Michaux C, Holmqvist E, Vasicek E, Sharan M, Barquist L, Westermann AJ, Gunn JS, Vogel J (2017)
PNAS 114 (26): 6824-6829
Resolving host-pathogen interactions by dual RNA-seq
Westermann AJ, Barquist L, Vogel J (2017)
PLOS Pathogens 13 (2): e1006033
2016
An expanded evaluation of protein function prediction methods shows an improvement in accuracy
Jiang Y, Oron TR, Clark WT, Bankapur AR, D'Andrea D, Lepore R, Funk CS, Kahanda I, Verspoor KM, Ben-Hur A, …, Friedberg I, Radivojac P (2016)
Genome Biology 17 (1): 184
The target spectrum of SdsR small RNA in Salmonella
Fröhlich KS, Haneke K, Papenfort K, Vogel J (2016)
Nucleic Acids Research 44 (21): 10406-22
A 3' UTR-Derived Small RNA Provides the Regulatory Noncoding Arm of the Inner Membrane Stress Response
Chao Y, Vogel J (2016)
Molecular Cell 61 (3): 352-363
Single-cell RNA-seq ties macrophage polarization to growth rate of intracellular Salmonella
Saliba AE, Li L, Westermann AJ, Appenzeller S, Stapels DA, Schulte LN, Helaine S, Vogel J (2016)
Nature Microbiology 2: 16206
Dual RNA-seq unveils noncoding RNA functions in host-pathogen interactions
Westermann AJ, Forstner KU, Amman F, Barquist L, Chao Y, Schulte LN, Muller L, Reinhardt R, Stadler PF, Vogel J (2016)
Nature 529 (7587): 496-501
cis-Encoded Small RNAs, a Conserved Mechanism for Repression of Polysaccharide Utilization in Bacteroides
Cao Y, Förstner KU, Vogel J, Smith CJ (2016)
Journal of Bacteriology 198 (18): 2410-8
The primary transcriptome of the Escherichia coli O104:H4 pAA plasmid and novel insights into its virulence gene expression and regulation
Berger P, Knödler M, Förstner KU, Berger M, Bertling C, Sharma CM, Vogel J, Karch H, Dobrindt U, Mellmann A (2016)
Scientific Reports 6: 35307
Natural mutations in a Staphylococcus aureus virulence regulator attenuate cytotoxicity but permit bacteremia and abscess formation
Das S, Lindemann C, Young BC, Muller J, Österreich B, Ternette N, Winkler A, Paprotka K, Reinhardt R, Förstner KU, …, Wyllie DH, Fraunholz MJ (2016)
PNAS 113 (22): E3101-10
An NK Cell Perforin Response Elicited via IL-18 Controls Mucosal Inflammation Kinetics during Salmonella Gut Infection
Müller AA, Dolowschiak T, Sellin ME, Felmy B, Verbree C, Gadient S, Westermann AJ, Vogel J, LeibundGut-Landmann S, Hardt W (2016)
PLOS Pathogens 12 (6): e1005723
Gifsy-1 Prophage IsrK with Dual Function as Small and Messenger RNA Modulates Vital Bacterial Machineries
Hershko-Shalev T, Odenheimer-Bergman A, Elgrably-Weiss M, Ben-Zvi T, Govindarajan S, Seri H, Papenfort K, Vogel J, Altuvia S (2016)
PLOS Genetics 12 (4): e1005975
Emerging roles of RNA modifications in bacteria
Marbaniang CN, Vogel J (2016)
Current Opinion in Microbiology 30: 50-57
Grad-seq guides the discovery of ProQ as a major small RNA-binding protein
Smirnov A, Förstner KU, Holmqvist E, Otto A, Günster R, Becher D, Reinhardt R, Vogel J (2016)
PNAS 113 (41): 11591-11596
Global RNA recognition patterns of post-transcriptional regulators Hfq and CsrA revealed by UV crosslinking in vivo
Holmqvist E, Wright PR, Li L, Bischler T, Barquist L, Reinhardt R, Backofen R, Vogel J (2016)
The EMBO Journal 35 (9): 991-1011
Molecular phenotyping of infection-associated small non-coding RNAs
Barquist L, Westermann AJ, Vogel J (2016)
Philosophical Transactions of the Royal Society of London B: Biological Sciences 371 (1707): 20160081
2015
The end is not the end: remnants of tRNA precursors live on to sponge up small regulatory RNAs
Ziebuhr W, Vogel J (2015)
Molecular Cell 58 (3): 389-90
Accelerating Discovery and Functional Analysis of Small RNAs with New Technologies
Barquist L, Vogel J (2015)
Annual Review of Genetics 49: 367-94
Small RNA-based feedforward loop with AND-gate logic regulates extrachromosomal DNA transfer in Salmonella
Papenfort K, Espinosa E, Casadesús J, Vogel J (2015)
PNAS 112 (34): E4772-81
dRNA-Seq Reveals Genomewide TSSs and Noncoding RNAs of Plant Beneficial Rhizobacterium Bacillus amyloliquefaciens FZB42
Fan B, Li L, Chao Y, Förstner K, Vogel J, Borriss R, Wu X (2015)
PLOS One 10 (11): e0142002
Cross talk between ABC transporter mRNAs via a target mRNA-derived sponge of the GcvB small RNA
Miyakoshi M, Chao Y, Vogel J (2015)
The EMBO Journal 34 (11): 1478-92
Investigating CRISPR RNA Biogenesis and Function Using RNA-seq
Heidrich N, Dugar G, Vogel J, Sharma CM (2015)
In: CRISPR: Methods and protocols (eds Lundgren M, Charpentier E, Fineran PC), Methods Mol Biol (Methods in Molecular Biology) 1311: 1-21
Regulatory small RNAs from the 3' regions of bacterial mRNAs
Miyakoshi M, Chao Y, Vogel J (2015)
Current Opinion in Microbiology 24: 132-9
Genome-wide transcription start site profiling in biofilm-grown Burkholderia cenocepacia J2315
Sass AM, van Acker H, Förstner KU, van Nieuwerburgh F, Deforce D, Vogel J, Coenye T (2015)
BMC Genomics 16: 775
Dual 3'Seq using deepSuperSAGE uncovers transcriptomes of interacting Salmonella enterica Typhimurium and human host cells
Afonso-Grunz F, Hoffmeier K, Müller S, Westermann AJ, Rotter B, Vogel J, Winter P, Kahl G (2015)
BMC Genomics 16: 323
2014
Identification and Characterization of Small Non-coding RNAs in Bacteria
Podkaminski D, Bouvier M, Vogel J (2014)
136: 719-786
READemption-a tool for the computational analysis of deep-sequencing-based transcriptome data
Förstner KU, Vogel J, Sharma CM (2014)
Bioinformatics 30 (23): 3421-3
Meeting report: Regulating with RNA in Bacteria 2013
Vogel J, Gottesman S, Belasco J, Narberhaus F (2014)
RNA Biology 11 (5): 403-12
Small RNA functions in carbon metabolism and virulence of enteric pathogens
Papenfort K, Vogel J (2014)
Frontiers in Cellular and Infection Microbiology 4: 91
Recognition of the small regulatory RNA RydC by the bacterial Hfq protein
Dimastrogiovanni D, Fröhlich KS, Bandyra KJ, Bruce HA, Hohensee S, Vogel J, Luisi BF (2014)
eLife 3: e05375
Differential RNA-seq: the approach behind and the biological insight gained
Sharma CM, Vogel J (2014)
Current Opinion in Microbiology 19: 97-105
Single-cell RNA-seq: advances and future challenges
Saliba AE, Westermann AJ, Gorski SA, Vogel J (2014)
Nucleic Acids Research 42 (14): 8845-60
2013
Same same but different: new structural insight into CRISPR-Cas complexes
Heidrich N, Vogel J (2013)
Molecular Cell 52 (1): 4-7
β-Lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity
Gutierrez A, Laureti L, Crussard S, Abida H, Rodríguez-Rojas A, Blázquez J, Baharoglu Z, Mazel D, Darfeuille F, Vogel J, Matic I (2013)
Nature Communications 4: 1610
Processing-Independent CRISPR RNAs Limit Natural Transformation in Neisseria meningitidis
Zhang Y, Heidrich N, Ampattu BJ, Gunderson CW, Seifert HS, Schoen C, Vogel J, Sontheimer EJ (2013)
Molecular Cell 50 (4): 488-503
Targeted decay of a regulatory small RNA by an adaptor protein for RNase E and counteraction by an anti-adaptor RNA
Göpel Y, Papenfort K, Reichenbach B, Vogel J, Görke B (2013)
Genes & Development 27 (5): 552-64
A small RNA activates CFA synthase by isoform-specific mRNA stabilization
Fröhlich KS, Papenfort K, Fekete A, Vogel J (2013)
The EMBO Journal 32 (22): 2963-79
CRISPRs extending their reach: prokaryotic RNAi protein Cas9 recruited for gene regulation
Heidrich N, Vogel J (2013)
The EMBO Journal 32 (13): 1802-4
Small RNA-Mediated Activation of Sugar Phosphatase mRNA Regulates Glucose Homeostasis
Papenfort K, Sun Y, Miyakoshi M, Vanderpool CK, Vogel J (2013)
Cell 153 (2): 426-437
A small RNA serving both the Hfq and CsrA regulons
Holmqvist E, Vogel J (2013)
Genes & Development 27 (10): 1073-8
Regulatory Mechanisms of Special Significance: Role of Small RNAs in Virulence Regulation
Papenfort K, Corcoran CP, Gupta SK, Miyakoshi M, Heidrich N, Chao Y, Fröhlich KS, Ziebuhr W, Böhm A, Vogel J, Sharma CM (2013)
In: Regulation of Bacterial Virulence (eds Darwin AJ, Vasil ML): 493-527
RNA-mediated regulation in pathogenic bacteria
Caldelari I, Chao Y, Romby P, Vogel J (2013)
Cold Spring Harbor Perspectives In Medicine 3 (9): a010298
Bacterial regulatory mechanisms: the gene and beyond
Bassler B, Vogel J (2013)
Current Opinion in Microbiology 16 (2): 109-11
Comparative genomics boosts target prediction for bacterial small RNAs
Wright PR, Richter AS, Papenfort K, Mann M, Vogel J, Hess WR, Backofen R, Georg J (2013)
PNAS 110 (37): E3487-96
Alexander Böhm (1971-2012)
Boos W, Parkinson JS, Jenal U, Vogel J, Søgaard-Andersen L (2013)
Molecular Microbiology 88 (2): 219-21
Differential activation and functional specialization of miR-146 and miR-155 in innate immune sensing
Schulte LN, Westermann AJ, Vogel J (2013)
Nucleic Acids Research 41 (1): 542-53
2012
The seed region of a small RNA drives the controlled destruction of the target mRNA by the endoribonuclease RNase E
Bandyra KJ, Said N, Pfeiffer V, Górna MW, Vogel J, Luisi BF (2012)
Molecular Cell 47 (6): 943-53
Dual RNA-seq of pathogen and host
Westermann AJ, Gorski SA, Vogel J (2012)
Nature Reviews Microbiology 10 (9): 618-30
The mammalian microRNA response to bacterial infections
Eulalio A, Schulte LN, Vogel J (2012)
RNA Biology 9 (6): 742-50
Experimental tools to identify RNA-protein interactions in Helicobacter pylori
Rieder R, Reinhardt R, Sharma CM, Vogel J (2012)
RNA Biology 9 (4): 520-31
Small RNAs of the Bradyrhizobium/Rhodopseudomonas lineage and their analysis
Madhugiri R, Pessi G, Voss B, Hahn J, Sharma CM, Reinhardt R, Vogel J, Hess WR, Fischer H, Evguenieva-Hackenberg E (2012)
RNA Biology 9 (1): 47-58
The ancestral SgrS RNA discriminates horizontally acquired Salmonella mRNAs through a single G-U wobble pair
Papenfort K, Podkaminski D, Hinton JC, Vogel J (2012)
PNAS 109 (13): E757-64
The transcriptional landscape and small RNAs of Salmonella enterica serovar Typhimurium
Kröger C, Dillon SC, Cameron AD, Papenfort K, Sivasankaran SK, Hokamp K, Chao Y, Sittka A, Hébrard M, Händler K, …, Vogel J, Hinton JC (2012)
PNAS 109 (20): E1277-86
RelA protein stimulates the activity of RyhB small RNA by acting on RNA-binding protein Hfq
Argaman L, Elgrably-Weiss M, Hershko T, Vogel J, Altuvia S (2012)
PNAS 109 (12): 4621-6
Global regulatory functions of the Staphylococcus aureus endoribonuclease III in gene expression
Lioliou E, Sharma CM, Caldelari I, Helfer A, Fechter P, Vandenesch F, Vogel J, Romby P (2012)
PLOS Biology 8 (6): e1002782
The Primary Transcriptome of Barley Chloroplasts: Numerous Noncoding RNAs and the Dominating Role of the Plastid-Encoded RNA Polymerase
Zhelyazkova P, Sharma CM, Förstner KU, Liere K, Vogel J, Börner T (2012)
The Plant Cell 24 (1): 123-36
Superfolder GFP reporters validate diverse new mRNA targets of the classic porin regulator, MicF RNA
Corcoran CP, Podkaminski D, Papenfort K, Urban JH, Hinton JC, Vogel J (2012)
Molecular Microbiology 84 (3): 428-45
The csgD mRNA as a hub for signal integration via multiple small RNAs
Boehm A, Vogel J (2012)
Molecular Microbiology 84 (1): 1-5
An atlas of Hfq-bound transcripts reveals 3' UTRs as a genomic reservoir of regulatory small RNAs
Chao Y, Papenfort K, Reinhardt R, Sharma CM, Vogel J (2012)
The EMBO Journal 31 (20): 4005-19
Use of aptamer tagging to identify in vivo protein binding partners of small regulatory RNAs
Corcoran CP, Rieder R, Podkaminski D, Hofmann B, Vogel J (2012)
In: Bacterial Regulatory RNA: Methods and protocols (ed Keiler KC) 905: 177-200
Hfq-associated Regulatory Small RNAs
Corcoran CP, Papenfort K, Vogel J (2012)
In: Regulatory RNAs in Prokaryotes (eds Marchfelder A, Hess WR): 15-50
Differential RNA Sequencing (dRNA-Seq): Deep-Sequencing-Based Analysis of Primary Transcriptomes
Borries A, Vogel J, Sharma CM (2012)
In: Tag-based next generation sequencing (eds Kahl G, Harbers M ) 316: 109-121
2011
Regulation by small RNAs in bacteria: expanding frontiers
Storz G, Vogel J, Wassarman KM (2011)
Molecular Cell 43 (6): 880-91
Sweet Business: Spot42 RNA networks with CRP to Modulate Catabolite Repression
Papenfort K, Vogel J (2011)
Molecular Cell 41 (3): 245-6
CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III
Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, Eckert MR, Vogel J, Charpentier E (2011)
Nature 471 (7340): 602-7
A conserved RpoS-dependent small RNA controls the synthesis of major porin OmpD
Fröhlich KS, Papenfort K, Berger AA, Vogel J (2011)
Nucleic Acids Research 40 (8): 3623-40
Helicobacter pylori interferes with an embryonic stem cell micro RNA cluster to block cell cycle progression
Belair C, Baud J, Chabas S, Sharma CM, Vogel J, Staedel C, Darfeuille F (2011)
Silence 2 (1): 7
Small RNAs endow a transcriptional activator with essential repressor functions for single-tier control of a global stress regulon
Gogol EB, Rhodius VA, Papenfort K, Vogel J, Gross CA (2011)
PNAS 108 (31): 12875-80
An experimentally anchored map of transcriptional start sites in the model cyanobacterium Synechocystis sp. PCC6803
Mitschke J, Georg J, Scholz I, Sharma CM, Dienst D, Bantscheff J, Voss B, Steglich C, Wilde A, Vogel J, Hess WR (2011)
PNAS 108 (5): 2124-9
A candidate approach implicates the secreted Salmonella effector protein SpvB in P-body disassembly
Eulalio A, Fröhlich KS, Mano M, Giacca M, Vogel J (2011)
PLOS One 6 (3): e17296
Genome-wide transcriptome analysis of the plant pathogen Xanthomonas identifies sRNAs with putative virulence functions
Schmidtke C, Findeiß S, Sharma CM, Kuhfuß J, Hoffmann S, Vogel J, Stadler PF, Bonas U (2011)
Nucleic Acids Research 40 (5): 2020-31
Pervasive post-transcriptional control of genes involved in amino acid metabolism by the Hfq-dependent GcvB small RNA
Sharma CM, Papenfort K, Pernitzsch SR, Mollenkopf H, Hinton JC, Vogel J (2011)
Molecular Microbiology 81 (5): 1144-65
Contribution of Hfq to photooxidative stress resistance and global regulation in Rhodobacter sphaeroides
Berghoff BA, Glaeser J, Sharma CM, Zobawa M, Lottspeich F, Vogel J, Klug G (2011)
Molecular Microbiology 80 (6): 1479-95
Microbes at their best: first Mol Micro Meeting Würzburg
Böhm A, Papenfort K, Lopez D, Vogel J (2011)
Molecular Microbiology 82 (4): 797-806
The transcriptional landscape of Chlamydia pneumoniae
Albrecht M, Sharma CM, Dittrich MT, Müller T, Reinhardt R, Vogel J, Rudel T (2011)
Genome Biology 12 (10): R98
Analysis of the host microRNA response to Salmonella uncovers the control of major cytokines by the let-7 family
Schulte LN, Eulalio A, Mollenkopf H, Reinhardt R, Vogel J (2011)
The EMBO Journal 30 (10): 1977-89
2010
The primary transcriptome of the major human pathogen Helicobacter pylori
Sharma CM, Hoffmann S, Darfeuille F, Reignier J, Findeiss S, Sittka A, Chabas S, Reiche K, Hackermüller J, Reinhardt R, Stadler PF, Vogel J (2010)
Nature 464 (7286): 250-5
The small RNAs of Salmonella
Javayel S, Papenfort K, Vogel J (2010)
In: Salmonella: From Genome to Function (ed Prowollik S)
Analysis of A to I editing of miRNA in macrophages exposed to salmonella
Heale BS, Eulalio A, Schulte LN, Vogel J, O'Connell MA (2010)
RNA Biology 7 (5): 621-7
Evidence for an autonomous 5′ target recognition domain in an Hfq-associated small RNA
Papenfort K, Bouvier M, Mika F, Sharma CM, Vogel J (2010)
PNAS 107 (47): 20435-40
Identification of regulatory RNAs in Bacillus subtilis
Irnov I, Sharma CM, Vogel J, Winkler WC (2010)
Nucleic Acids Research 38 (19): 6637-51
Deep sequencing-based discovery of the Chlamydia trachomatis transcriptome
Albrecht M, Sharma CM, Reinhardt R, Vogel J, Rudel T (2010)
Nucleic Acids Research 38 (3): 868-77
Experimental discovery of small RNAs in Staphylococcus aureus reveals a riboregulator of central metabolism
Bohn C, Rigoulay C, Chabelskaya S, Sharma CM, Marchais A, Skorski P, Borezée-Durant E, Barbet R, Jacquet E, Jacq A, …, Vogel J, Bouloc P (2010)
Nucleic Acids Research 38 (19): 6620-36
Small RNAs promote mRNA stability to activate the synthesis of virulence factors
Podkaminski D, Vogel J (2010)
Molecular Microbiology 78 (6): 1327-31
The role of Hfq in bacterial pathogens
Chao Y, Vogel J (2010)
Current Opinion in Microbiology 13 (1): 24-33
2009
Coding sequence targeting by MicC RNA reveals bacterial mRNA silencing downstream of translational initiation
Pfeiffer V, Papenfort K, Lucchini S, Hinton JC, Vogel J (2009)
Nature Structural & Molecular Biology 16 (8): 840-6
A small RNA cascade regulates aminosugar synthesis
Vogel J (2009)
In: Nova Acta Leopoldina 378: 41-50
A green fluorescent protein (GFP)-based plasmid system to study post-transcriptional control of gene expression in vivo
Urban JH, Vogel J (2009)
In: Riboswitches: Methods and protocols (ed Serganov A), Methods Mol Biol 540: 301-19
Deep sequencing of Salmonella RNA associated with heterologous Hfq proteins in vivo reveals small RNAs as a major target class and identifies RNA processing phenotypes
Sittka A, Sharma CM, Rolle K, Vogel J (2009)
RNA Biology 6 (3): 266-75
Multiple target regulation by small noncoding RNAs rewires gene expression at the post-transcriptional level
Papenfort K, Vogel J (2009)
Research in Microbiology 160 (4): 278-87
Deep sequencing analysis of the Methanosarcina mazei Gö1 transcriptome in response to nitrogen availability
Jäger D, Sharma CM, Thomsen J, Ehlers C, Vogel J, Schmitz RA (2009)
PNAS 106 (51): 21878-82
In vivo expression and purification of aptamer-tagged small RNA regulators
Said N, Rieder R, Hurwitz R, Deckert J, Urlaub H, Vogel J (2009)
Nucleic Acids Research 37 (20): e133-e133
An RNA trap helps bacteria get the most out of chitosugars
Vogel J (2009)
Molecular Microbiology 73 (5): 737-41
A rough guide to the non-coding RNA world of Salmonella
Vogel J (2009)
Molecular Microbiology 71 (1): 1-11
Regulatory RNAs in prokaryotes: here, there and everywhere
Narberhaus F, Vogel J (2009)
Molecular Microbiology 74 (2): 261-9
Photooxidative stress-induced and abundant small RNAs in Rhodobacter sphaeroides
Berghoff BA, Glaeser J, Sharma CM, Vogel J, Klug G (2009)
Molecular Microbiology 74 (6): 1497-512
Specific and pleiotropic patterns of mRNA regulation by ArcZ, a conserved, Hfq-dependent small RNA
Papenfort K, Said N, Welsink T, Lucchini S, Hinton JC, Vogel J (2009)
Molecular Microbiology 74 (1): 139-158
Acid stress activation of the sigma(E) stress response in Salmonella enterica serovar Typhimurium
Muller C, Bang I, Velayudhan J, Karlinsey J, Papenfort K, Vogel J, Fang FC (2009)
Molecular Microbiology 71 (5): 1228-38
Fast mapping of short sequences with mismatches, insertions and deletions using index structures
Hoffmann S, Otto C, Kurtz S, Sharma CM, Khaitovich P, Vogel J, Stadler PF, Hackermüller J (2009)
PLOS Computational Biology 5 (9): e1000502
Experimental approaches for the discovery and characterization of regulatory small RNA
Sharma CM, Vogel J (2009)
Current Opinion in Microbiology 12 (5): 536-46
Activation of gene expression by small RNA
Fröhlich KS, Vogel J (2009)
Current Opinion in Microbiology 12 (6): 674-82
2008
Two seemingly homologous noncoding RNAs act hierarchically to activate glmS mRNA translation
Urban JH, Vogel J (2008)
PLOS Biology 6 (3): e64
Small RNA binding to 5' mRNA coding region inhibits translational initiation
Bouvier M, Sharma CM, Mika F, Nierhaus KH, Vogel J (2008)
Molecular Cell 32 (6): 827-837
Noncoding RNA control of the making and breaking of sugars
Görke B, Vogel J (2008)
Genes & Development 22 (21): 2914-25
The cyanobacterial homologue of the RNA chaperone Hfq is essential for motility of Synechocystis sp. PCC 6803
Dienst D, Dühring U, Mollenkopf H, Vogel J, Golecki J, Hess WR, Wilde A (2008)
Microbiology 154 (Pt 10): 3134-3143
RNA: Noncoding
Henry AA, Vogel J (2008)
In: Wiley Encyclopedia of Chemical Biology (ed Begley TP) 22: 2183
Deep sequencing analysis of small noncoding RNA and mRNA targets of the global post-transcriptional regulator, Hfq
Sittka A, Lucchini S, Papenfort K, Sharma CM, Rolle K, Binnewies TT, Hinton JC, Vogel J (2008)
PLOS Genetics 4 (8): e1000163
A new Vibrio cholerae sRNA modulates colonization and affects release of outer membrane vesicles
Song T, Mika F, Lindmark B, Liu Z, Schild S, Bishop A, Zhu J, Camilli A, Johansson J, Vogel J, Wai SN (2008)
Molecular Microbiology 70 (1): 100-111
Systematic deletion of Salmonella small RNA genes identifies CyaR, a conserved CRP-dependent riboregulator of OmpX synthesis
Papenfort K, Pfeiffer V, Lucchini S, Sonawane A, Hinton JC, Vogel J (2008)
Molecular Microbiology 68 (4): 890-906
A Glimpse at the Evolution of Virulence Control
Sittka A, Vogel J (2008)
Cell Host & Microbe 4 (4): 310-312
2007
An antisense RNA inhibits translation by competing with standby ribosomes
Darfeuille F, Unoson C, Vogel J, Wagner EG (2007)
Molecular Cell 26 (3): 381-92
A small RNA regulates multiple ABC transporter mRNAs by targeting C/A-rich elements inside and upstream of ribosome-binding sites
Sharma CM, Darfeuille F, Plantinga TH, Vogel J (2007)
Genes & Development 21 (21): 2804-17
Recent advances in the expression, evolution, and dynamics of prokaryotic genomes
Arraiano CM, Bamford J, Brüssow H, Carpousis AJ, Pelicic V, Pflüger K, Polard P, Vogel J (2007)
Journal of Bacteriology 189 (17): 6093-100
Sensory and regulatory RNAs in prokaryotes: a new German research focus
Narberhaus F, Vogel J (2007)
RNA Biology 4 (3): 160-4
The RNA chaperone Hfq is essential for the virulence of Salmonella typhimurium
Sittka A, Pfeiffer V, Tedin K, Vogel J (2007)
Molecular Microbiology 63 (1): 193-217
Translational control and target recognition by Escherichia coli small RNAs in vivo
Urban JH, Vogel J (2007)
Journal of Molecular Biology 35 (3): 1018-37
A conserved small RNA promotes discoordinate expression of the glmUS operon mRNA to activate GlmS synthesis
Urban JH, Papenfort K, Thomsen J, Schmitz RA, Vogel J (2007)
Journal of Molecular Biology 373 (3): 521-8
Characterization of the role of ribonucleases in Salmonella small RNA decay
Viegas SC, Pfeiffer V, Sittka A, Silva IJ, Vogel J, Arraiano CM (2007)
Nucleic Acids Research 35 (22): 7651-64
Space flight alters bacterial gene expression and virulence and reveals a role for global regulator Hfq
Wilson JW, Ott CM, Höner zu Bentrup K, Ramamurthy R, Quick L, Porwollik S, Cheng P, McClelland M, Tsaprailis G, Radabaugh T, …, Stefanyshyn-Piper HM, Nickerson CA (2007)
PNAS 104 (41): 16299-304
A small non-coding RNA of the invasion gene island (SPI-1) represses outer membrane protein synthesis from the Salmonella core genome
Pfeiffer V, Sittka A, Tomer R, Tedin K, Brinkmann V, Vogel J (2007)
Molecular Microbiology 66 (5): 1174-91
Target identification of small noncoding RNAs in bacteria
Vogel J, Wagner EG (2007)
Current Opinion in Microbiology 10 (3): 262-70
2006
Small non-coding RNAs and the bacterial outer membrane
Vogel J, Papenfort K (2006)
Current Opinion in Microbiology 9 (6): 605-11
Experimental approaches to identify non-coding RNAs
Hüttenhofer A, Vogel J (2006)
Nucleic Acids Research 34 (2): 635-46
σE-dependent small RNAs of Salmonella respond to membrane stress by accelerating global omp mRNA decay
Papenfort K, Pfeiffer V, Mika F, Lucchini S, Hinton JC, Vogel J (2006)
Molecular Microbiology 62 (6): 1674-1688
2005
Hfq-dependent regulation of OmpA synthesis is mediated by an antisense RNA
Udekwu KI, Darfeuille F, Vogel J, Reimegård J, Holmqvist E, Wagner EG (2005)
Genes & Development 19 (19): 2355-66
Approaches to Identify Novel Non-messenger RNAs in Bacteria and to Investigate their Biological Functions: Functional Analysis of Identified Non-mRNAs
Gerhart E, Wagner H, Vogel J (2005)
In: Handbook of RNA Biochemistry (eds Hartmann RK, Bindereif A, Schön A, Westhof E) 2: 614-642
Approaches to Identify Novel Non-messenger RNAs in Bacteria and to Investigate their Biological Functions: RNA Mining
Vogel J, Gerhart E, Wagner H (2005)
In: Handbook of RNA Biochemistry (eds Hartmann RK, Bindereif A, Schön A, Westhof E) 2: 595-613
How to find small non-coding RNAs in bacteria
Vogel J, Sharma CM (2005)
Biological Chemistry 386 (12): 1219-38
Identification of cyanobacterial non-coding RNAs by comparative genome analysis
Axmann IM, Kensche P, Vogel J, Kohl S, Herzel H, Hess WR (2005)
Genome Biology 6 (9): R73
2004
The small RNA IstR inhibits synthesis of an SOS-induced toxic peptide
Vogel J, Argaman L, Wagner EG, Altuvia S (2004)
Current Biology 14 (24): 2271-6
2003
Noncoding RNAs encoded by bacterial chromosomes
Wagner EG, Vogel J (2003)
In: Noncoding RNAs (eds Barciszewski J, Erdmann V): 243-259
Experimental and computational analysis of transcriptional start sites in the cyanobacterium Prochlorococcus MED4
Vogel J, Axmann IM, Herzel H, Hess WR (2003)
Nucleic Acids Research 31 (11): 2890-9
RNomics in Escherichia coli detects new sRNA species and indicates parallel transcriptional output in bacteria
Vogel J, Bartels V, Tang TH, Churakov G, Slagter-Jäger JG, Hüttenhofer A, Wagner EG (2003)
Nucleic Acids Research 31 (22): 6435-43
2002
Lariat formation and a hydrolytic pathway in plant chloroplast group II intron splicing
Vogel J, Börner T (2002)
The EMBO Journal 21 (14): 3794-803
2001
Complete 5' and 3' end maturation of group II intron-containing tRNA precursors
Vogel J, Hess WR (2001)
RNA 7 (2): 285-92
Novel small RNA-encoding genes in the intergenic regions of Escherichia coli
Argaman L, Hershberg R, Vogel J, Bejerano G, Wagner EG, Margalit H, Altuvia S (2001)
Current Biology 11 (12): 941-50
The ins and outs of group II introns
Bonen L, Vogel J (2001)
Trends In Genetics 17 (6): 322-31