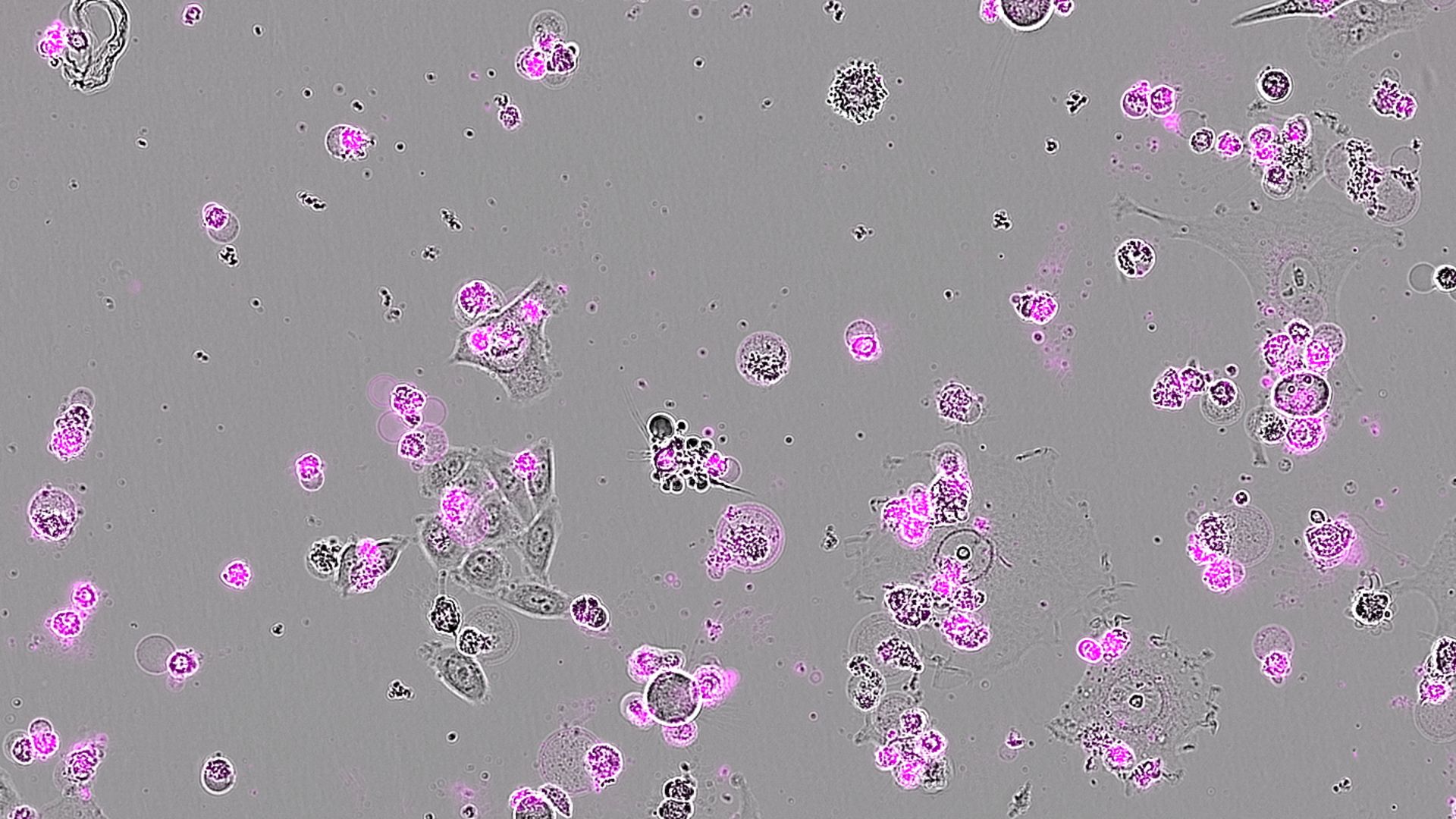
Discover Our Institute
People & Research at HIRI
Upcoming Conferences & Events
Research Groups at HIRI

RNA Biology of Bacterial Infections
Prof Jörg Vogel

Single-cell Analysis
Prof Antoine-Emmanuel Saliba

Molecular Principles of RNA Phages
Jun Prof Jens Hör

Systems Microbiology of Intracellular Pathogens
Jun Prof Camilla Ciolli Mattioli

RNA Biology of gram-positive Bacteria (associated research group)
Prof Franziska Faber
Associated & Alumni Group Leaders

RNA Synthetic Biology
Dr Chase Beisel, Affiliated Department Head

Recoding Mechanisms in Infections
Prof Neva Caliskan, Former Group Leader

Integrative Informatics for Infection Biology
Prof Lars Barquist, Associated Scientist

LncRNA and Infection Biology
Prof Mathias Munschauer, Former Group Leader

Host-pathogen-microbiota interactions
Prof Alexander Westermann, Affiliated Scientist

Genome Architecture and Evolution of RNA viruses
Dr Redmond Smyth, Former Group Leader
HIRI at a Glance
HZI + JMU = HIRI
- Foundation May 2017.
- Site of the Braunschweig Helmholtz Centre for Infection Research (HZI) in cooperation with the Julius-Maximilians-Universität Würzburg (JMU).
Managing Director
- Prof Jörg Vogel, Director since 2017.
- Prof Vogel is also Director and Full Professor (W3) at the Institute for Molecular Infection Biology, JMU Würzburg.
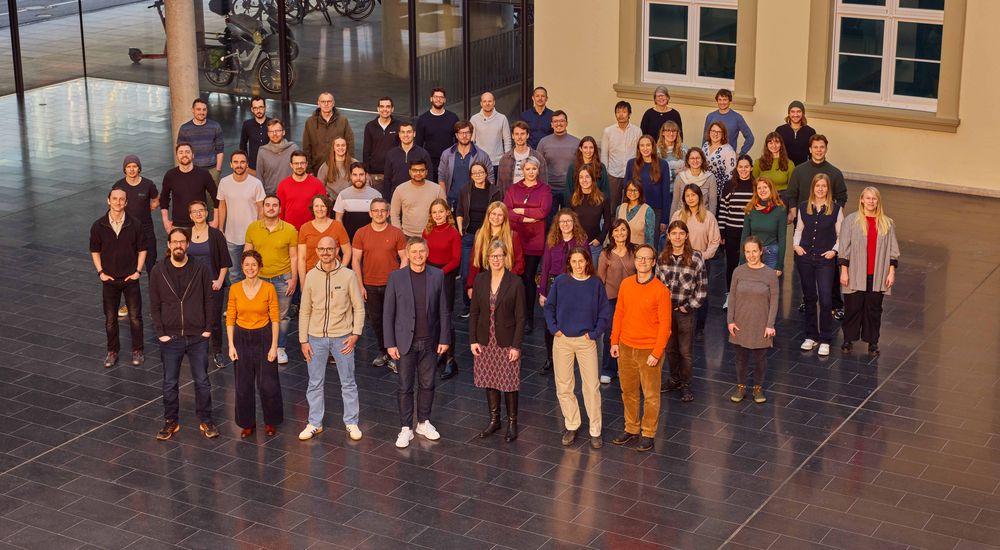
Team
- 5 research groups.
- 6 associated & alumni group leaders.
Funding
- Sound annual core funding from federal government and Free State of Bavaria.
- Third party funding rate of 30%.
- 7 ERC Grants.
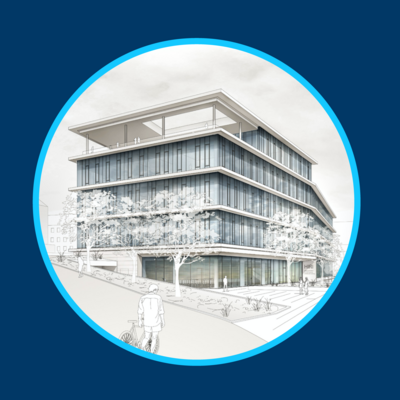
New Building
- State-of-the-art facilities.
- Funded by Free State of Bavaria, co-financed by European Union.
- Completion planned for 2027.
Profile Areas & Programs
- Single-Cell Center Würzburg.
- Graduate Training Programs “RNA & Infection” and “RNAmed - Future Leaders in RNA-based Medicine”.
More about the HIRI
“RNA is a powerful medium for engineering biology. We seek to better understand the fundamental properties of this biomolecule and how it can be harnessed to improve diagnosis and treatment of human infections.”
Chase Beisel

GET IN TOUCH
If you would like to inquire about job offers, ask scientific questions or simply pay us a visit, you find all necessary information on this page. We look forward to your email, call or visit!