Dr Chase Beisel, Affiliated Department Head
RNA Synthetic Biology
In 2018, the Helmholtz Institute Würzburg (HIRI) established a research focus on CRISPR biology and technologies. Group leader Chase Beisel, who was appointed in 2025 as a faculty member at the Botnar Institute of Immune Engineering (BIIE) in Basel (Switzerland), directs the research with the overarching goal of developing new technologies that can be employed to better understand, diagnose and treat human infections.
Our research and approach
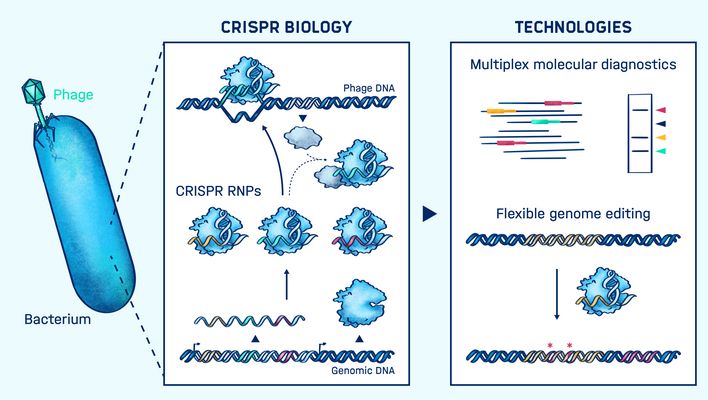
The discovery of CRISPR-Cas immune systems in microbes has led to the development of revolutionary tools for genome editing and many other applications benefiting society. In infection biology, CRISPR tools can be used to interrogate pathogens, microbiota and human cells as well as to diagnose and treat infections.
Chase Beisel’s group adopts synthetic biology approaches to investigate how these immune systems function and how they can be converted into different technologies. They interrogate the properties of the immune systems using cell-free transcription-translation systems as well as E. coli and non-model bacteria naturally harboring these systems. They also regularly employ high-throughput approaches such as next-generation sequencing and library screening to collect extensive information from individual experiments.
The group has developed and advanced a number of important CRISPR-based technologies, including the use of CRISPR for genome editing and gene regulation in bacteria, for multi-test diagnostics, and for tailored-spectrum antimicrobials. Their overarching goal is to exploit CRISPR technologies to advance our understanding of human infections and develop diagnostic tools together with tailored therapeutic approaches.
Team members

Dr Chase Beisel, Affiliated Department Head
Group Leader

Dr Akanksha Roberts
Postdoc

Thomas Gaudin
PhD Student

Tatjana Achmedov
Technical Assistant
Research projects
Graphical Abstract
Introducing the research group
Alumni
Adini Arifah, PhD Student • Harris Bassett, PhD Student • Irene Calvin, Technical Assistant • Xiye Chen, PhD Student • Daphne Collias, Postdoc • Alessandro Del Re, PhD Student • Oleg Dmytrenko, Postdoc • Lilly Erkenbrecher, Volunteer (FSJ) • Sandra Gawlitt, PhD Student • Mohammad Ghaem-Maghami, Postdoc • Darshana Gupta, PhD Student • Jimmy Jeske, Technical Assistant • Chunlei Jiao, Postdoc • Charlotte Kamm, PhD Student • Anuja Kibe, Postdoc • Isabell Köblitz, Technical Assistent • Rachael Larose, Technical Assistant • Chunyu Liao, Postdoc • Sudeshna Manna, Postdoc • Angela Migur, Postdoc • Ioannis Mougiakos, Postdoc • Ahsen Özcan, Postdoc • Costas Patinios, Postdoc • Friso Schut, PhD Student • Elena Vialetto, PhD Student • Katharina Wandera, PhD Student • Yanyan Wang, Technical Assistant • Franziska Wimmer, PhD Student • Yan Yan, Technical Assistant • Jiaqi Yu, Technical Assistant
Publications
2026
RNA-triggered cell killing with CRISPR-Cas12a2
Scholz P, Thompson J, Crosby KT, Fauth T, Krah NM, Schlauderaff G, Back R, Berkheimer ZA, Jolley A, Sombroek D, …, Beisel CL, Liu Y (2026)
Nature (Online ahead of print)
A leader-repeat hairpin blocks extraneous CRISPR RNA production in diverse CRISPR-Cas13 systems
Migur A, Feussner M, Liao C, Alkhnbashi OS, Chauvier A, Walter NG, Backofen R, Weinberg Z, Beisel CL, Migur A, …, Weinberg Z, Beisel CL (2026)
The EMBO Journal 45 (10): 3396-3415
Comprehensive analysis of CRISPR array repeat mutations reveals subtype-specific patterns and links to spacer dynamics
Mitrofanov A, Beisel CL, Baumdicker F, Alkhnbashi OS, Backofen R (2026)
microLife 7: uqaf050
2025
Streamlined and efficient genome editing in Cupriavidus necator H16 using an optimised SIBR-Cas system
Della Valle S, Orsi E, Creutzburg SCA, Jansen LFM, Pentari EN, Beisel CL, Steel H, Nikel PI, Staals RHJ, Claassens NJ, …, Huang WE, Patinios C (2025)
Trends in Biotechnology 43 (6): 1470-1491
AcrVIB1 inhibits CRISPR-Cas13b immunity by promoting unproductive crRNA binding accessible to RNase attack
Wandera KG, Schmelz S, Migur A, Kibe A, Lukat P, Achmedov T, Caliskan N, Blankenfeldt W, Beisel CL (2025)
Molecular Cell 85 (6): 1162-1175
The translational landscape of HIV-1 infected cells reveals key gene regulatory principles
Kibe A, Buck S, Gribling-Burrer AS, Gilmer O, Bohn P, Koch T, Mireisz CN, Schlosser A, Erhard F, Smyth RP, Caliskan N (2025)
Nature Structural & Molecular Biology 32 (5): 841-852
A resurrected ancestor of Cas12a expands target access and substrate recognition for nucleic acid editing and detection
Jabalera Y, Tascón I, Samperio S, López-Alonso JP, Gonzalez-Lopez M, Aransay AM, Abascal-Palacios G, Beisel CL, Ubarretxena-Belandia I, Perez-Jimenez R (2025)
Nature Biotechnology 43 (10): 1663-1672
An updated evolutionary classification of CRISPR-Cas systems including rare variants
Makarova KS, Shmakov SA, Wolf YI, Mutz P, Altae-Tran H, Beisel CL, Brouns SJJ, Charpentier E, Cheng D, Doudna J, …, Barrangou R, Koonin EV (2025)
Nature Microbiology 10 (12): 3346-3361
Targeted DNA ADP-ribosylation triggers templated repair in bacteria and base mutagenesis in eukaryotes
Gupta D, Patinios C, Bassett HV, Kibe A, Collins SP, Kamm C, Wang Y, Zhao C, Vollen K, Toussaint C, …, Schut F, Corn JE (2025)
Nature Biotechnology (Online ahead of print)
PAM-interacting domain turn-helix 51 motifs can improve Cas9-SpRY activity
Eggenschwiler R, Hoffmann T, Dmytrenko O, Opitz M, Ackel-Zakour M, Wang P, McCallan SA, Fráguas-Eggenschwiler M, Niemann H, Patronov A, Beisel CL, Cantz T (2025)
Nucleic Acids Research 53 (15): gkaf782
Cas9-independent tracrRNA cytotoxicity in Lacticaseibacillus paracasei
Arifah AQ, Vento JM, Kurrer I, Achmedov T, Beisel CL (2025)
microLife 6: uqaf013
Efficient encapsulation of CRISPR-Cas9 RNP in bioreducible nanogels and release in a cytosol-mimicking environment
Westarp P, Keller T, Brand J, Horvat S, Albrecht K, Beisel C, Groll J (2025)
Discover Nano 20 (1): 119
Disparate mechanisms counteract extraneous CRISPR RNA production in type II-C CRISPR-Cas systems
Feussner M, Migur A, Mitrofanov A, Alkhnbashi OS, Backofen R, Beisel CL, Weinberg Z (2025)
microLife 6: uqaf007
Purification and in vivo, cell-free, and in vitro characterization of CRISPR-Cas12a2
Schut FT, Hallmark T, Dmytrenko O, Jackson RN, Beisel CL (2025)
Methods In Enzymology 712: 143-181
2024
Phage anti-CRISPR control by an RNA- and DNA-binding helix-turn-helix protein
Birkholz N, Kamata K, Feussner M, Wilkinson ME, Cuba Samaniego C, Migur A, Kimanius D, Ceelen M, Went SC, Usher B, …, Jackson SA, Fineran PC (2024)
Nature 631 (8021): 670-677
TracrRNA reprogramming enables direct PAM-independent detection of RNA with diverse DNA-targeting Cas12 nucleases
Jiao C, Peeck NL, Yu J, Ghaem Maghami M, Kono S, Collias D, Martinez Diaz SL, Larose R, Beisel CL (2024)
Nature Communications 15 (1): 5909
A cell-free transcription-translation pipeline for recreating methylation patterns boosts DNA transformation in bacteria
Vento JM, Durmusoglu D, Li T, Patinios C, Sullivan S, Ttofali F, van Schaik J, Yu Y, Wang Y, Barquist L, Crook N, Beisel CL (2024)
Molecular Cell 84 (14): 2785-2796.e4
Type IV-A3 CRISPR-Cas systems drive inter-plasmid conflicts by acquiring spacers in trans
Benz F, Camara-Wilpert S, Russel J, Wandera KG, Cepaite R, Ares-Arroyo M, Gomes-Filho JV, Englert F, Kuehn JA, Gloor S, …, Sørensen SJ, Pinilla-Redondo R (2024)
Cell Host & Microbe 32 (6): 875-886.e9
Expanding the flexibility of base editing for high-throughput genetic screens in bacteria
Gawlitt S, Collins SP, Yu Y, Blackman SA, Barquist L, Beisel CL (2024)
Nucleic Acids Research 52 (7): 4079-4097
CRISPR-based screening of small RNA modulators of bile susceptibility in Bacteroides thetaiotaomicron
Prezza G, Liao C, Reichardt S, Beisel CL, Westermann AJ (2024)
PNAS 121 (6): 1096
Improved prediction of bacterial CRISPRi guide efficiency from depletion screens through mixed-effect machine learning and data integration
Yu Y, Gawlitt S, de Andrade E Sousa LB, Merdivan E, Piraud M, Beisel CL, Barquist L (2024)
Genome Biology 25 (1): 13
Phages to the rescue: in situ editing of the gut microbiota
Kamm C, Beisel CL (2024)
Trends in Microbiology 32 (10): 934-935
Cell-free expression with a quartz crystal microbalance enables rapid, dynamic, and label-free characterization of membrane-interacting proteins
Khakimzhan A, Izri Z, Thompson S, Dmytrenko O, Fischer P, Beisel C, Noireaux V (2024)
Communications Biology 7 (1): 1005
MprF-mediated immune evasion is necessary for Lactiplantibacillus plantarum resilience in the Drosophila gut during inflammation
Arias-Rojas A, Arifah AQ, Angelidou G, Alshaar B, Schombel U, Forest E, Frahm D, Brinkmann V, Paczia N, Beisel CL, Gisch N, Iatsenko I (2024)
PLOS Pathogens 20 (8): e1012462
A Hitchhiker's guide to CRISPR editing tools in bacteria : CRISPR can help unlock the bacterial world, but technical and regulatory barriers persist
Krink N, Nikel PI, Beisel CL (2024)
EMBO Reports 25 (4): 1694-1699
An adapted method for Cas9-mediated editing reveals the species-specific role of β-glucoside utilization driving competition between Klebsiella species
Almási ÉdH, Knischewski N, Osbelt L, Muthukumarasamy U, El Mouali Y, Vialetto E, Beisel CL, Strowig T (2024)
Journal of Bacteriology 206 (3): e0031723
2023
Interrogating two extensively self-targeting Type I CRISPR-Cas systems in Xanthomonas albilineans reveals distinct anti-CRISPR proteins that block DNA degradation
Wimmer F, Englert F, Wandera KG, Alkhnbashi OS, Collins SP, Backofen R, Beisel CL (2023)
Nucleic Acids Research 52 (2): 769-783
A predicted CRISPR-mediated symbiosis between uncultivated archaea
Esser SP, Rahlff J, Zhao W, Predl M, Plewka J, Sures K, Wimmer F, Lee J, Adam PS, McGonigle J, …, Zhang Y, Probst AJ (2023)
Nature Microbiology 8 (9): 1619-1633
Systematically attenuating DNA targeting enables CRISPR-driven editing in bacteria
Collias D, Vialetto E, Yu J, Co K, Almási ÉDH, Rüttiger AS, Achmedov T, Strowig T, Beisel CL (2023)
Nature Communications 14 (1): 680
RNA targeting unleashes indiscriminate nuclease activity of CRISPR-Cas12a2
Bravo JPK, Hallmark T, Naegle B, Beisel CL, Jackson RN, Taylor DW (2023)
Nature 613 (7944): 582-587
Cas12a2 elicits abortive infection through RNA-triggered destruction of dsDNA
Dmytrenko O, Neumann GC, Hallmark T, Keiser DJ, Crowley VM, Vialetto E, Mougiakos I, Wandera KG, Domgaard H, Weber J, …, Jackson RN, Beisel CL (2023)
Nature 613 (7944): 588-594
RNA recording in single bacterial cells using reprogrammed tracrRNAs
Jiao C, Reckstadt C, König F, Homberger C, Yu J, Vogel J, Westermann AJ, Sharma CM, Beisel CL (2023)
Nature Biotechnology 41 (8): 1107-1116
SND1 binds SARS-CoV-2 negative-sense RNA and promotes viral RNA synthesis through NSP9
Schmidt N, Ganskih S, Wei Y, Gabel A, Zielinski S, Keshishian H, Lareau CA, Zimmermann L, Makroczyova J, Pearce C, …, Erhard F, Munschauer M (2023)
Cell 186 (22): 4834-4850.e23
Shortened CRISPR-Cas9 arrays enable multiplexed gene targeting in bacteria from a smaller DNA footprint
Gawlitt S, Liao C, Achmedov T, Beisel CL (2023)
RNA Biology 20 (1): 666-680
For the CRISPR Fan(zor)atics: RNA-guided DNA endonucleases discovered in eukaryotes
Patinios C, Beisel CL (2023)
Molecular Cell 83 (17): 3046-3048
Optimized metrics for orthogonal combinatorial CRISPR screens
Cetin R, Wegner M, Luwisch L, Saud S, Achmedov T, Süsser S, Vera-Guapi A, Müller K, Matthess Y, Quandt E, …, Beisel CL, Kaulich M (2023)
Scientific Reports 13 (1): 7405
2022
The miniature CRISPR-Cas12m effector binds DNA to block transcription
Wu WY, Mohanraju P, Liao C, Adiego-Pérez B, Creutzburg SCA, Makarova KS, Keessen K, Lindeboom TA, Khan TS, Prinsen S, …, Beisel CL, van der Oost J (2022)
Molecular Cell 82 (23): 4487-4502.e7
Anti-CRISPR prediction using deep learning reveals an inhibitor of Cas13b nucleases
Wandera KG, Alkhnbashi OS, Bassett HVI, Mitrofanov A, Hauns S, Migur A, Backofen R, Beisel CL (2022)
Molecular Cell 82 (14): 2714-2726.e4
A target expression threshold dictates invader defense and prevents autoimmunity by CRISPR-Cas13
Vialetto E, Yu Y, Collins SP, Wandera KG, Barquist L, Beisel CL (2022)
Cell Host & Microbe 3128 (22): 00273-6
Rapid cell-free characterization of multi-subunit CRISPR effectors and transposons
Wimmer F, Mougiakos I, Englert F, Beisel CL (2022)
Molecular Cell 82 (6): 1210-1224.e6
Spacer prioritization in CRISPR-Cas9 immunity is enabled by the leader RNA
Liao C, Sharma S, Svensson SL, Kibe A, Weinberg Z, Alkhnbashi OS, Bischler T, Backofen R, Caliskan N, Sharma CM, Beisel CL (2022)
Nature Microbiology 7 (4): 530-541
Cell-Free Protein Synthesis from Exonuclease-Deficient Cellular Extracts Utilizing Linear DNA Templates
Sabeti Azad M, Cardoso Batista A, Faulon JL, Beisel CL, Bonnet J, Kushwaha M (2022)
Journal of Visualized Experiments (186): e64236
Reprogramming TracrRNAs for Multiplexed RNA Detection
Jiao C, Beisel CL (2022)
Methods in Molecular Biology 2518: 217-235
Beneficial commensal bacteria promote Drosophila growth by downregulating the expression of peptidoglycan recognition proteins
Gallo M, Vento JM, Joncour P, Quagliariello A, Maritan E, Silva-Soares NF, Battistolli M, Beisel CL, Martino ME (2022)
iScience 25 (6): 104357
Genome Editing with Cas9 in Lactobacilli
Vento JM BC (2022)
Methods in Molecular Biology 2479: 245-261
Rapidly Characterizing CRISPR-Cas13 Nucleases Using Cell-Free Transcription-Translation Systems
Wandera KG, Beisel CL (2022)
Methods in Molecular Biology 2404: 135-153
Differentially Optimized Cell-Free Buffer Enables Robust Expression from Unprotected Linear DNA in Exonuclease-Deficient Extracts
Batista AC, Levrier A, Soudier P, Voyvodic PL, Achmedov T, Reif-Trauttmansdorff T, DeVisch A, Cohen-Gonsaud M, Faulon JL, Beisel CL, Bonnet J, Kushwaha M (2022)
ACS Synthetic Biology 11 (2): 732-746
A TXTL-Based Assay to Rapidly Identify PAMs for CRISPR-Cas Systems with Multi-Protein Effector Complexes
Wimmer F, Englert F, Beisel CL (2022)
Methods in Molecular Biology 2433: 391-411
2021
The tracrRNA in CRISPR Biology and Technologies
Liao C, Beisel CL (2021)
Annual Review of Genetics 55: 161-181
A genetically encoded anti-CRISPR protein constrains gene drive spread and prevents population suppression
Taxiarchi C, Beaghton A, Don NI, Kyrou K, Gribble M, Shittu D, Collins SP, Beisel CL, Galizi R, Crisanti A (2021)
Nature Communications 12 (1): 3977
Noncanonical crRNAs derived from host transcripts enable multiplexable RNA detection by Cas9
Jiao C, Sharma S, Dugar G, Peeck NL, Bischler T, Wimmer F, Yu Y, Barquist L, Schoen C, Kurzai O, Sharma CM, Beisel CL (2021)
Science 372 (6545): 941-948
Sequence-independent RNA sensing and DNA targeting by a split domain CRISPR-Cas12a gRNA switch
Collins SP, Rostain W, Liao C, Beisel CL (2021)
Nucleic Acids Research 49 (5): 2985-2999
CRISPR technologies and the search for the PAM-free nuclease
Collias D, Beisel CL (2021)
Nature Communications 12 (1): 555
Coupling smartphone and CRISPR–Cas12a for digital and multiplexed nucleic acid detection
Yu T, Zhang S, Matei R, Marx W, Beisel CL, Wei Q (2021)
American Institute of Chemical Engineers Journal 67 (12): 17399‐17405
A Hyperthermoactive-Cas9 Editing Tool Reveals the Role of a Unique Arsenite Methyltransferase in the Arsenic Resistance System of Thermus thermophilus HB27
Gallo G, Mougiakos I, Bianco M, Carbonaro M, Carpentieri A, Illiano A, Pucci P, Bartolucci S, van der Oost J, Fiorentino G (2021)
mBio 12 (6): e0281321
In Situ Biomanufacturing of Small Molecules in the Mammalian Gut by Probiotic Saccharomyces boulardii
Durmusoglu D, Al'Abri IS, Collins SP, Cheng J, Eroglu A, Beisel CL, Crook N (2021)
ACS Synthetic Biology 10 (5): 1039-1052
A Code of Ethics for Gene Drive Research
Annas GJ, Beisel CL, Clement K, Crisanti A, Francis S, Galardini M, Galizi R, Grünewald J, Immobile G, Khalil AS, …, Taxiarchi C, Joung JK (2021)
The CRISPR Journal 4 (1): 19-24
2020
A positive, growth-based PAM screen identifies noncanonical motifs recognized by the S. pyogenes Cas9
Collias D, Leenay RT, Slotkowski RA, Zuo Z, Collins SP, McGirr BA, Liu J, Beisel CL (2020)
Science Advances 6 (29): eabb4054
Your Base Editor Might Be Flirting with Single (Stranded) DNA: Faithful On-Target CRISPR Base Editing without Promiscuous Deamination
Collins SP, Beisel CL (2020)
Molecular Cell 79 (5): 703-704
Characterization of Cas12a nucleases reveals diverse PAM profiles between closely-related orthologs
Jacobsen T, Ttofali F, Liao C, Manchalu S, Gray BN, Beisel CL (2020)
Nucleic Acids Research 48 (10): 5624-5638
Rapid Testing of CRISPR Nucleases and Guide RNAs in an E. coli Cell-Free Transcription-Translation System
Marshall R, Beisel CL, Noireaux V (2020)
STAR Protocols 1 (1): 100003
Tunable self-cleaving ribozymes for modulating gene expression in eukaryotic systems
Jacobsen T, Yi G, Al Asafen H, Jermusyk AA, Beisel CL, Reeves GT (2020)
PLOS One 15 (4): e0232046
Competitive exclusion is a major bioprotective mechanism of lactobacilli against fungal spoilage in fermented milk products
Siedler S, Rau MH, Bidstrup S, Vento JM, Aunsbjerg SD, Bosma EF, McNair LM, Beisel CL, Neves AR (2020)
Applied and Environmental Microbiology 86 (7): e02312-19
Methods for characterizing, applying, and teaching CRISPR-Cas systems
Beisel CL (2020)
Methods 172: 1-2
CRISPR-Cas Systems and the Paradox of Self-Targeting Spacers
Wimmer F, Beisel CL (2020)
Frontiers in Microbiology 10: 3078
2019
Targeted transcriptional modulation with type I CRISPR-Cas systems in human cells
Pickar-Oliver A, Black JB, Lewis MM, Mutchnick KJ, Klann TS, Gilcrest KA, Sitton MJ, Nelson CE, Barrera A, Bartelt LC, …, Barrangou R, Gersbach CA (2019)
Nature Biotechnology 37 (12): 1493-1501
Modular one-pot assembly of CRISPR arrays enables library generation and reveals factors influencing crRNA biogenesis
Liao C, Ttofali F, Slotkowski RA, Denny SR, Cecil TD, Leenay RT, Keung AJ, Beisel CL (2019)
Nature Communications 10: 2948
An educational module to explore CRISPR technologies with a cell-free transcription-translation system
Collias D, Marshall R, Collins SP, Beisel CL, Noireaux V (2019)
Synthetic Biology 4 (1): 156
CRATES: A one-step assembly method for Class 2 CRISPR arrays
Liao C, Slotkowski RA, Beisel CL (2019)
Methods In Enzymology 629: 493-511
Barriers to genome editing with CRISPR in bacteria
Vento JM, Crook N, Beisel CL (2019)
Journal of Industrial Microbiology & Biotechnology 46 (9-10): 1327-1341
Characterization of the all-E. coli transcription-translation system myTXTL by mass spectrometry
Garenne D, Beisel CL, Noireaux V (2019)
Rapid Communications In Mass Spectrometry 33 (11): 1036-1048
Distinct timescales of RNA regulators enable the construction of a genetic pulse generator
Westbrook A, Tang X, Marshall R, Maxwell CS, Chappell J, Agrawal DK, Dunlop MJ, Noireaux V, Beisel CL, Lucks J, Franco E (2019)
Biotechnology and Bioengineering 116 (5): 1139-1151
An enhanced assay to characterize anti-CRISPR proteins using a cell-free transcription-translation system
Wandera KG, Collins SP, Wimmer F, Marshall R, Noireaux V, Beisel CL (2019)
Methods 172: 42-50
The Acidaminococcus sp. Cas12a nuclease recognizes GTTV and GCTV as non-canonical PAMs
Jacobsen T, Liao C, Beisel CL (2019)
FEMS Microbiology Letters 366 (8): pii: fnz085
2018
Bacterial Adaptation to the Host's Diet Is a Key Evolutionary Force Shaping Drosophila-Lactobacillus Symbiosis
Martino ME, Joncour P, Leenay R, Gervais H, Shah M, Hughes S, Gillet B, Beisel CL, Leulier F (2018)
Cell Host & Microbe 24 (1): 109-119.e6
Rapid and Scalable Characterization of CRISPR Technologies Using an E. coli Cell-Free Transcription-Translation System
Marshall R, Maxwell CS, Collins SP, Jacobsen T, Luo ML, Begemann MB, Gray BN, January E, Singer A, He Y, Beisel CL, Noireaux V (2018)
Molecular Cell 69 (1): 146-157.e3
CRISPR RNA-Dependent Binding and Cleavage of Endogenous RNAs by the Campylobacter jejuni Cas9
Dugar G, Leenay RT, Eisenbart SK, Bischler T, Aul BU, Beisel CL, Sharma CM (2018)
Molecular Cell 69 (5): 893-905.e7
Advances in CRISPR Technologies for Microbial Strain Engineering
Alper HS, Beisel CL (2018)
Biotechnology Journal 13 (9): e1800460
Synthetic Biology Approaches to Engineer Probiotics and Members of the Human Microbiota for Biomedical Applications
Bober JR, Beisel CL, Nair NU (2018)
Annual Review of Biomedical Engineering 20: 277-300
Mathematical Modeling of RNA-Based Architectures for Closed Loop Control of Gene Expression
Agrawal DK, Tang X, Westbrook A, Marshall R, Maxwell CS, Lucks J, Noireaux V, Beisel CL, Dunlop MJ, Franco E (2018)
ACS Synthetic Biology 7 (5): 1219-1228
A detailed cell-free transcription-translation-based assay to decipher CRISPR protospacer-adjacent motifs
Maxwell CS, Jacobsen T, Marshall R, Noireaux V, Beisel CL (2018)
Methods 143: 48-57
Genome Editing with CRISPR-Cas9 in Lactobacillus plantarum Revealed That Editing Outcomes Can Vary Across Strains and Between Methods
Leenay RT, Vento JM, Shah M, Martino ME, Leulier F, Beisel CL (2018)
Biotechnology Journal 14 (3): e1700583
The Francisella novicida Cas12a is sensitive to the structure downstream of the terminal repeat in CRISPR arrays
Liao C, Slotkowski RA, Achmedov T, Beisel CL (2018)
RNA Biology 16 (4): 404-412
2017
What Is the Role of Circuit Design in the Advancement of Synthetic Biology? Part 1
Beisel CL (2017)
Cell Systems 4 (4): 370-372
Deciphering, Communicating, and Engineering the CRISPR PAM
Leenay RT, Beisel CL (2017)
Journal of Molecular Biology 429 (2): 177-191
Toward a genetic tool development pipeline for host-associated bacteria
Waller MC, Bober JR, Nair NU, Beisel CL (2017)
Current Opinion in Microbiology 38: 156-164
Short DNA containing χ sites enhances DNA stability and gene expression in E. coli cell-free transcription-translation systems
Marshall R, Maxwell CS, Collins SP, Beisel CL, Noireaux V (2017)
Biotechnology and Bioengineering 114 (9): 2137-41
Advancing the design and delivery of CRISPR antimicrobials
Fagen JR, Collias D, Singh AK, Beisel CL (2017)
Current Opinion in Biomedical Engineering 4: 57-64
2016
The CRISPR RNA-guided surveillance complex in Escherichia coli accommodates extended RNA spacers
Luo ML, Jackson RN, Denny SR, Tokmina-Lukaszewska M, Maksimchuk KR, Lin W, Bothner B, Wiedenheft B, Beisel CL (2016)
Nucleic Acids Research 44 (15): 7385-94
Identifying and Visualizing Functional PAM Diversity across CRISPR-Cas Systems
Leenay RT, Maksimchuk KR, Slotkowski RA, Agrawal RN, Gomaa AA, Briner AE, Barrangou R, Beisel CL (2016)
Molecular Cell 62 (1): 137-47
SBE Supplement: Synthetic Biology – Engineering Genes with CRISPR-Cas9
Luo ML, Beisel CL (2016)
American Institute of Chemical Engineers
Synthetic Biology - Engineering Genes with CRISPR-Cas9
Luo ML, Beisel CL (2016)
CEP Magazine September
Rethinking the Hierarchy of Sugar Utilization in Bacteria
Beisel CL, Afroz T (2016)
Journal of Bacteriology 198 (3): 374-376
Current and future prospects for CRISPR-based tools in bacteria
Luo ML, Leenay RT, Beisel CL (2016)
Biotechnology and Bioengineering 113 (5): 930-43
2015
Self‐Assembled DNA Nanoclews for the Efficient Delivery of CRISPR–Cas9 for Genome Editing
Sun W, Ji W, Hall JM, Hu Q, Wang C, Beisel CL, Gu Z (2015)
Angewandte Chemie International Edition 54 (41): 12029-33
Integrative, ligand-responsive microRNAs
Smolke CD, Beisel CL (2015)
Patent (US9145555B2)
Trade-offs in engineering sugar utilization pathways for titratable control
Afroz T, Biliouris K, Boykin KE, Kaznessis Y, Beisel CL (2015)
ACS Synthetic Biology 4 (2): 141-9
Impact of Residual Inducer on Titratable Expression Systems
Afroz T, Luo ML, Beisel CL (2015)
PLOS One 10 (9): e0137421
Repurposing endogenous type I CRISPR-Cas systems for programmable gene repression
Luo ML, Mullis AS, Leenay RT, Beisel CL (2015)
Nucleic Acids Research 43 (1): 674-81
2014
A CRISPR design for next-generation antimicrobials
Beisel CL, Gomaa AA, Barrangou R (2014)
Genome Biology 15 (11): 516
Guide RNA functional modules direct Cas9 activity and orthogonality
Briner AE, Donohoue PD, Gomaa AA, Selle K, Slorach EM, Nye CH, Haurwitz RE, Beisel CL, May AP, Barrangou R (2014)
Molecular Cell 56 (2): 333-339
Construction of ligand-responsive microRNAs that operate through inhibition of Drosha processing
Beisel CL, Bloom RJ, Smolke CD (2014)
In: Artificial Riboswitches (ed Ogawa A), Methods Mol Biol 1111: 259-67
Bacterial sugar utilization gives rise to distinct single-cell behaviours
Afroz T, Biliouris K, Kaznessis Y, Beisel CL (2014)
Molecular Microbiology 93 (6): 1093-1103
Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems
Gomaa AA, Klumpe HE, Luo ML, Selle K, Barrangou R, Beisel CL (2014)
mBio 5 (1): e00928-13
2013
Modular polynucleotides for ligand-controlled regulatory systems
Smolke CD, Win MN, Beisel CL (2013)
Patent (US8367815B2)
Understanding and exploiting feedback in synthetic biology
Afroz T, Beisel CL (2013)
Chemical Engineering Science 103: 79-90
2012
Multiple factors dictate target selection by Hfq-binding small RNAs
Beisel CL, Updegrove TB, Janson BJ, Storz G (2012)
The EMBO Journal 31 (8): 1961-74
2011
The base pairing RNA Spot 42 participates in a multi-output feedforward loop to help enact catabolite repression in Escherichia coli
Beisel CL, Storz G (2011)
Molecular Cell 41 (3): 286-97
Design of small molecule-responsive microRNAs based on structural requirements for Drosha processing
Beisel CL, Chen YY, Culler SJ, Hoff KG, Smolke CD (2011)
Nucleic Acids Research 39 (7): 2981-94
Discriminating tastes: physiological contributions of the Hfq-binding small RNA Spot 42 to catabolite repression
Beisel CL, Storz G (2011)
RNA Biology 8 (5): 766-70
2010
Base pairing small RNAs and their roles in global regulatory networks
Beisel CL, Storz G (2010)
FEMS Microbiology Reviews 34 (5): 866-82
2009
Synthetic control of a fitness tradeoff in yeast nitrogen metabolism
Bayer TS, Hoff KG, Beisel CL, Lee JJ, Smolke CD (2009)
Journal of Biological Engineering 3: 1
Design Principles for Riboswitch Function
Beisel CL, Smolke CD (2009)
PLOS Computational Biology 5 (4): e1000363
2008
Model-guided design of ligand-regulated RNAi for programmable control of gene expression
Beisel CL, Bayer TS, Hoff KG, Smolke CD (2008)
Molecular Systems Biology 4: 224
2005
Conformational analysis of gossypol and its derivatives by molecular mechanics
Beisel CL, Dowd MK, Reilly PJ (2005)
Journal Of Molecular Structure: Computational and Theoretical Chemistry 730 (1-3): 51-58
2002
Cochlear whole mount in situ hybridization: identification of longitudinal and radial gradients
Judice TN, Nelson NC, Beisel CL, Delimont DC, Fritzsch B, Beisel KW (2002)
Brain Research Protocols 9 (1): 65-76