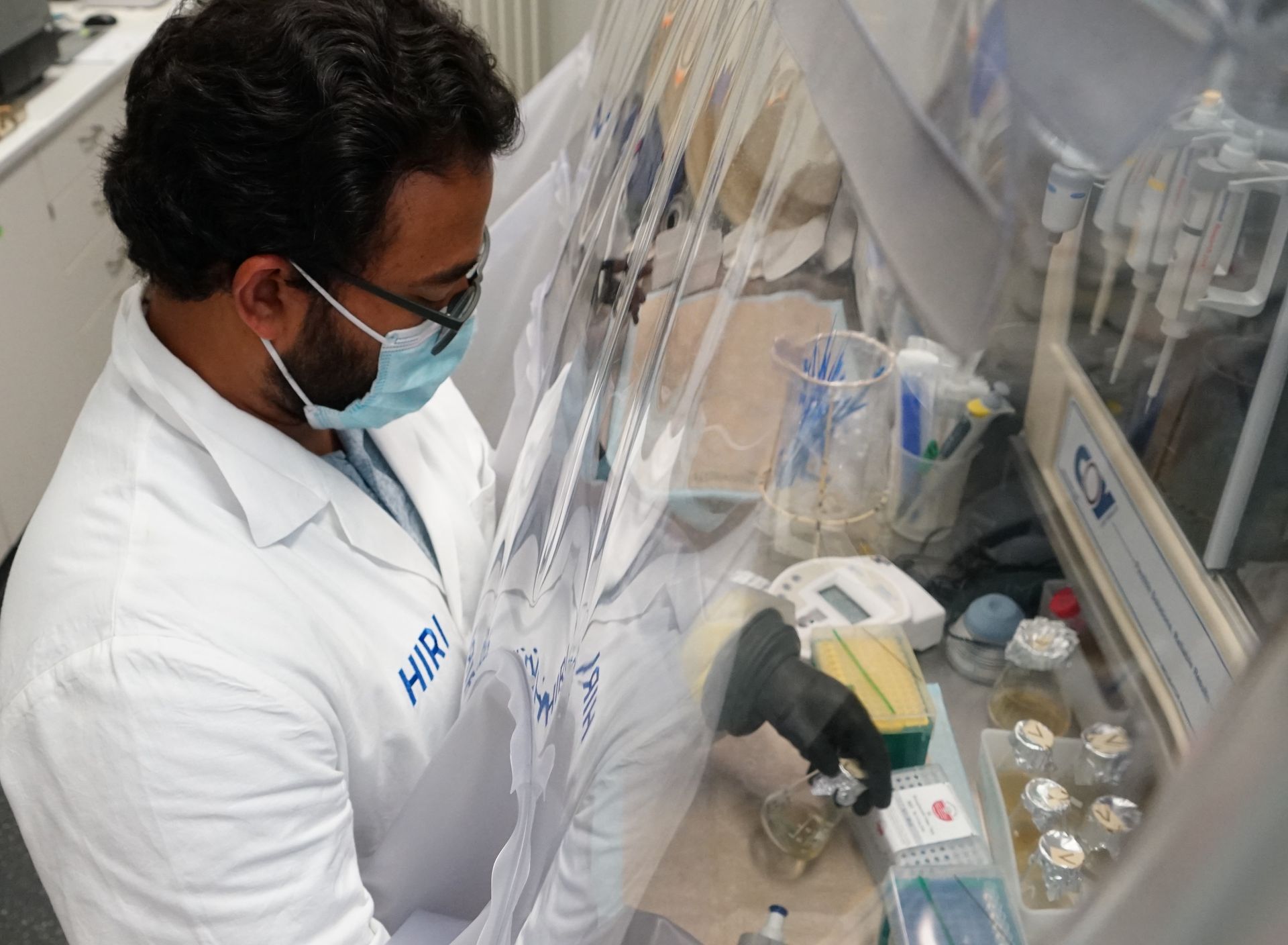
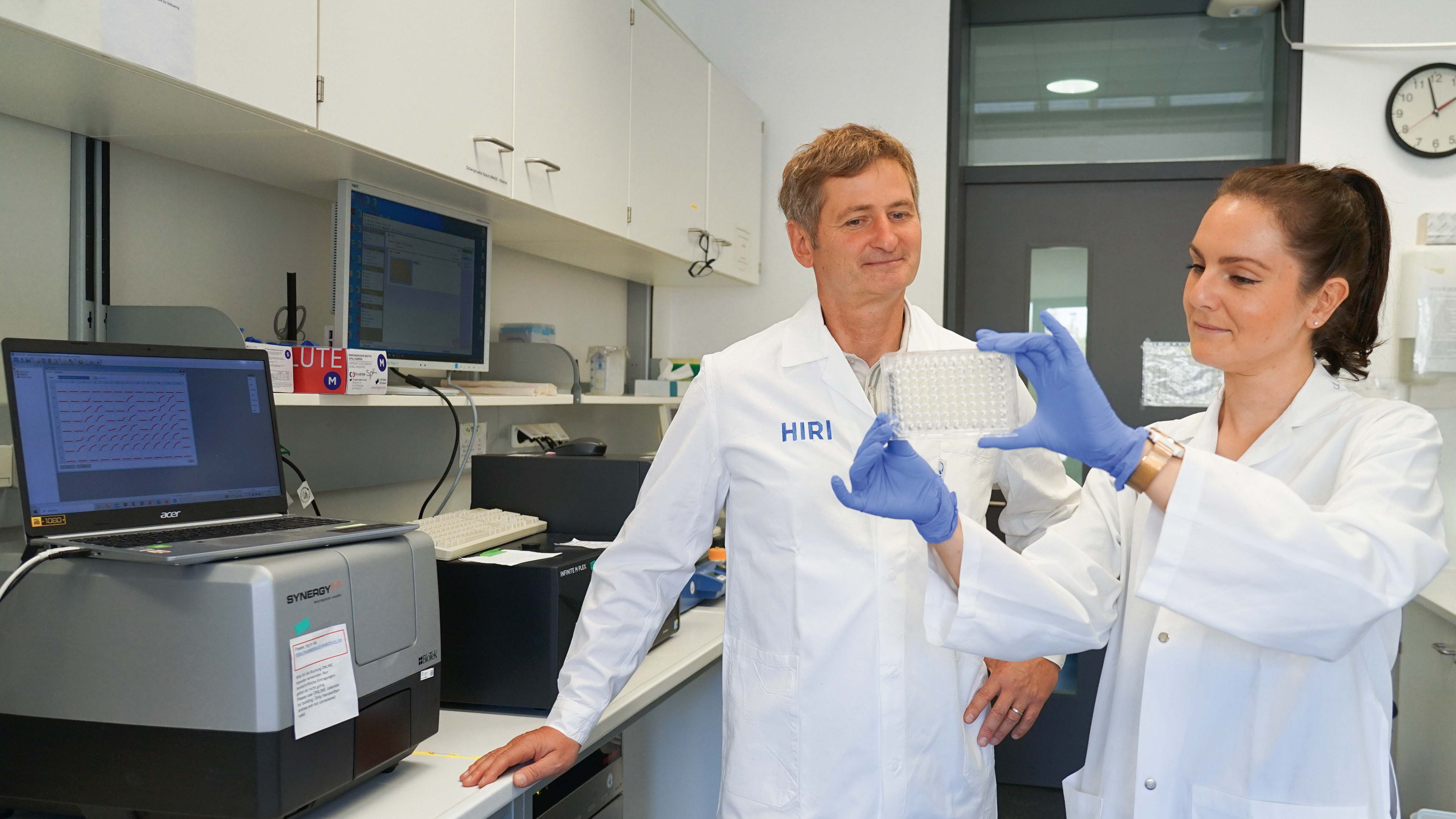
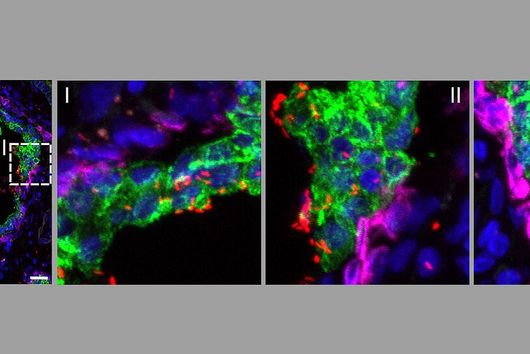
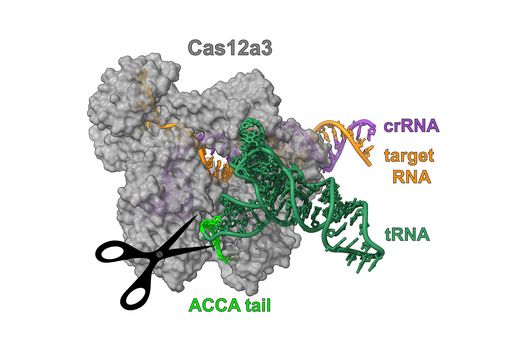
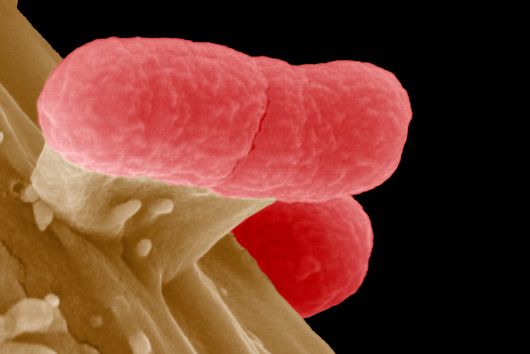
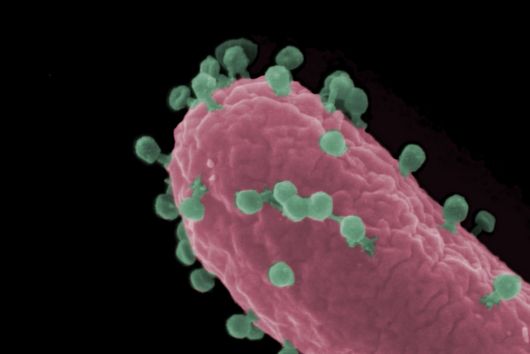
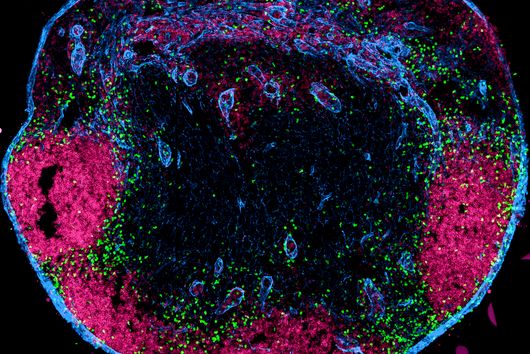
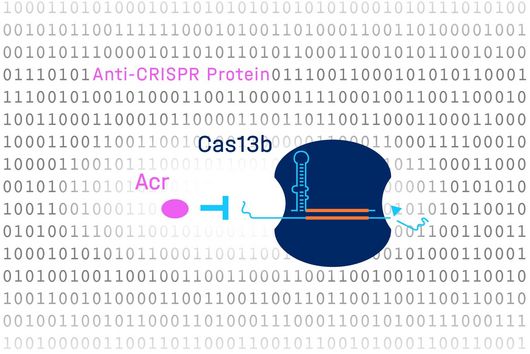
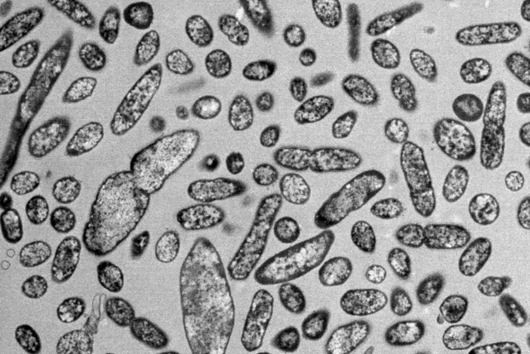
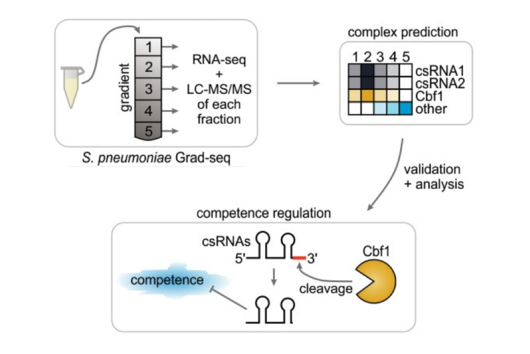
The intestinal microbiota plays a central role for human physiology. Traditional studies have focused on the changes in microbiota composition under diverse conditions. Recently, efforts shifted towards understanding the molecular biology of intestinal bacteria. The species Bacteroides thetaiotaomicron (B. theta) has emerged as a model organism for human intestinal microbiota research. Scientists at the Helmholtz Institute for RNA-based Infection Research (HIRI) have captured a detailed map of RNAs and their activities in these anaerobic bacteria. The HIRI is a joint institution of the Helmholtz Centre for Infection Research (HZI) in Braunschweig and the Julius-Maximilians-Universität Würzburg. The results can be browsed in an online database and were published in the journal Nature Communications.
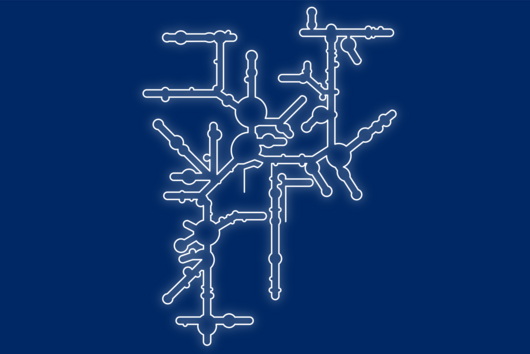
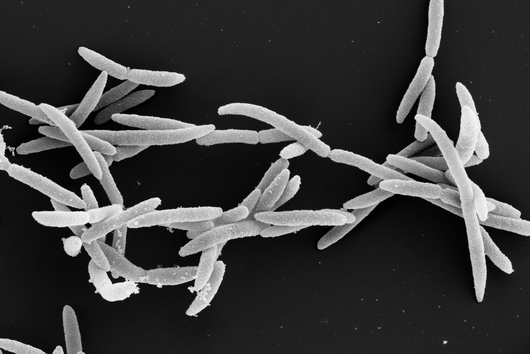
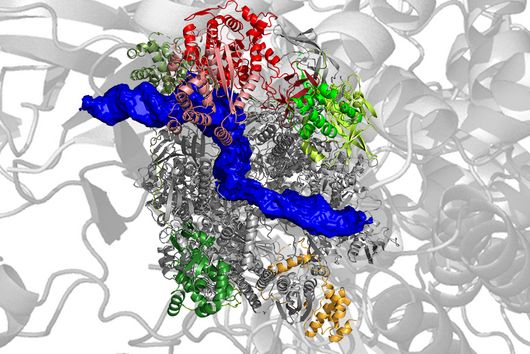
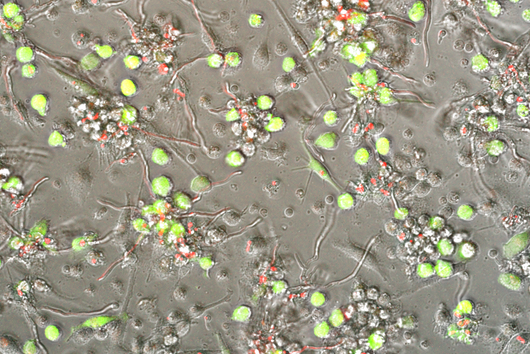
In the human intestine, countless bacteria form a complex community. B. theta is one of the most abundant species there and relatively easy to study in the lab. ”Until now, no fine-grained transcriptome annotation – a map of the entirety of RNA molecules – was available. Without this guide, it is very difficult to address basic questions of the organism’s biology,” says Jun Prof Alexander Westermann, head of the HIRI research group “Host-pathogen-microbiota interactions“. Using RNA sequencing, the Westermann group in close collaboration with Lars Barquist’s team captured all RNAs produced by B. theta. Their transcriptomic data are available to the research community in the online browser “Theta base” – a resource generated to facilitate research routines of many scientists working in this field.
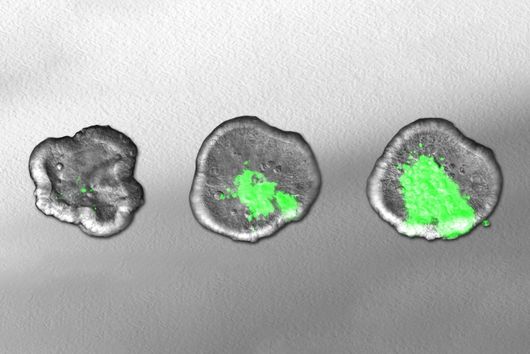
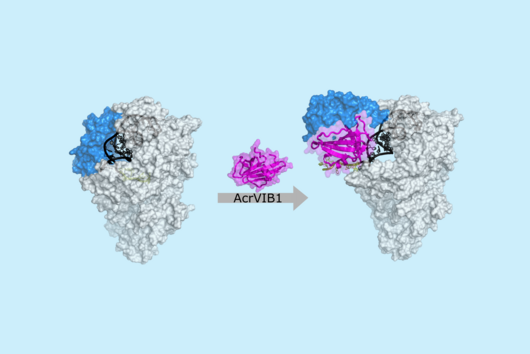
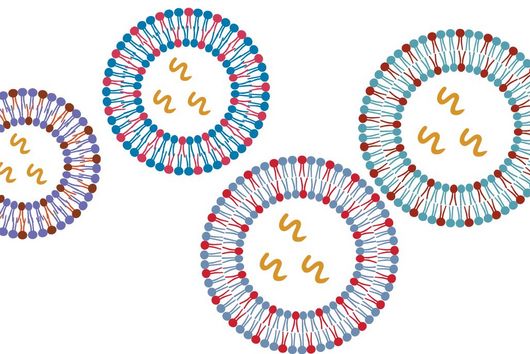
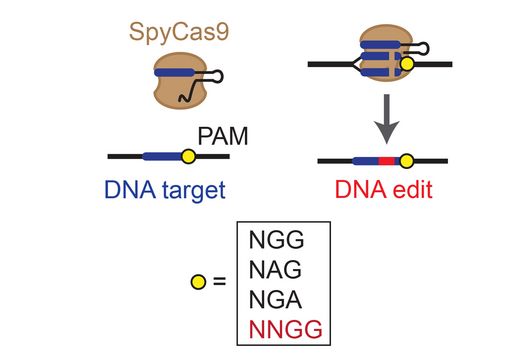
In the dataset, the HIRI researchers found more than 200 noncoding RNAs. Unlike messenger RNA that serves as a blueprint for proteins, noncoding RNA is not translated. Instead, many of them act as regulatory RNA, affecting the activity of target genes through riboregulation. B. theta – like all members of the Bacteroidetes family – lacks RNA-binding chaperone proteins common to other bacteria. “Therefore, it was unclear how widespread riboregulation actually is in Bacteroides. We found indications for ultra-conserved as well as highly specific RNA-based regulators,” says Westermann. In their study, the researchers characterised the conserved small RNA (sRNA) GibS. They found that specific sugar molecules trigger the production of this sRNA. When these monosaccharides are sensed by the bacteria, GibS modulates the expression of metabolic target genes. This may allow B. theta to rapidly adapt to changing nutrient availabilities. “Compared with aerobic bacteria, little is known about RNA-based gene regulation in anaerobic species – now we are finally starting to explore that world. Our next steps will be to determine how noncoding RNAs support B. theta’s lifestyle in the human intestine,” says Westermann.
Original publication
Daniel Ryan, Laura Jenniches, Sarah Reichardt, Lars Barquist, Alexander J. Westermann: A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron. Nature Communications 2020. DOI: 10.1038/s41467-020-17348-5
Funding
This work was financially supported by the DFG (Grant WE 6689/1-1) and by the IZKF at the University of Würzburg (project Z-6).