Decoding SARS-CoV-2
Researchers from the Helmholtz Institute Würzburg unveiled the interplay of RNA structures during the translation of the SARS-CoV-2 genome
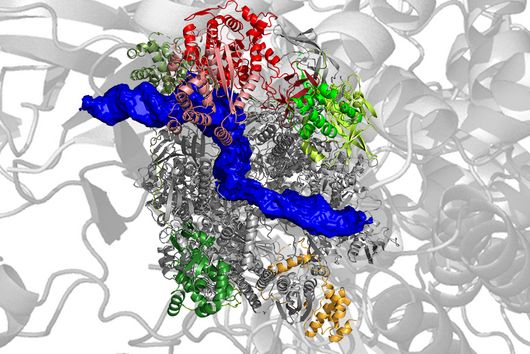
Würzburg, March 23, 2023—Translation of viral ribonucleic acids (RNAs) is often driven by non-canonical events, which are encoded in the viral genome in order to take over the control of the cellular translation machinery. One such non-canonical event is programmed ribosomal frameshifting. It prompts the translating ribosome to slip one nucleotide back or forth into a different reading frame, thus enabling viral replication. Using a variety of cutting-edge technologies, researchers from the Helmholtz Institute Würzburg (HIRI) unveiled the structural transitions in the SARS-CoV-2 frameshifting process. Their findings could lead to the discovery of new therapeutic targets and were published in the journal Nucleic Acids Research.
Translation is an essential process in every living cell and key to producing proteins. Viruses have developed a spectrum of non-canonical events at every step of translation. This way, they can efficiently hijack cellular protein synthesis. This is also the case for SARS-CoV-2, which uses the programmed ribosomal frameshifting to enable the expression of its replication enzymes.
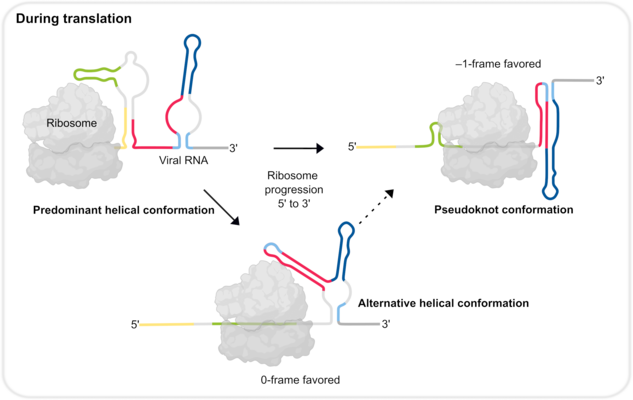
The efficiency of programmed ribosomal frameshifting depends on the presence of certain parts of the frameshift stimulatory element within the viral RNA. One of these parts is a stable structure that pauses translating ribosomes over slippery nucleotides allowing the ribosomes to slip into an alternative reading frame. In the case of SARS-CoV-2, this structure was originally proposed to form a pseudoknot, which represents a stable RNA fold acting as a roadblock for the ribosome.
However, recent attempts by other researchers to study the structure of the full-length viral genome failed to detect the pseudoknot structure being present. This apparent inconsistency in the literature has attracted the attention of scientists from the Helmholtz Institute for RNA-based Infection Research (HIRI) in Würzburg, a site of the Braunschweig Helmholtz Centre for Infection Research in cooperation with the Julius-Maximilians-Universität (JMU) of Würzburg.
Breaking down the virus
In the current study, which was published in the journal Nucleic Acids Research, a team led by HIRI group leader Neva Caliskan focused on the piece of viral RNA responsible for programmed ribosomal frameshifting. “Our aim was to acquire the complete picture of the frameshifting mechanism of SARS-CoV-2,” says Caliskan. Using cutting-edge technologies like single-molecule optical tweezers, the researchers studied a handful of viral RNA variants differing in length as well as some mutations. This way, they were able to analyze the folding of RNA structures in real-time. To complement the structural information, a functional assay was employed to determine the frameshifting efficiency. In a parallel approach to confirm the single-molecule results, the lab of HIRI group leader Redmond Smyth investigated the RNA structures by a chemical probing method.
The orchestrated interplay of RNA structures
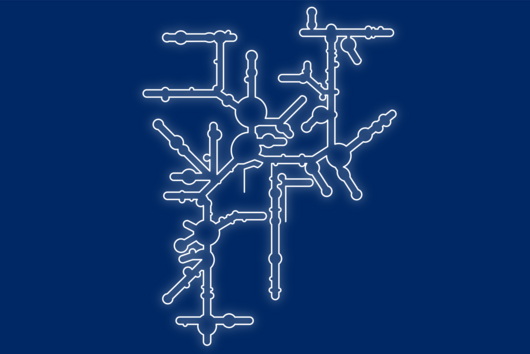
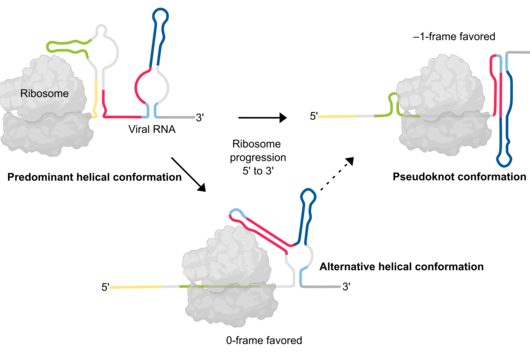
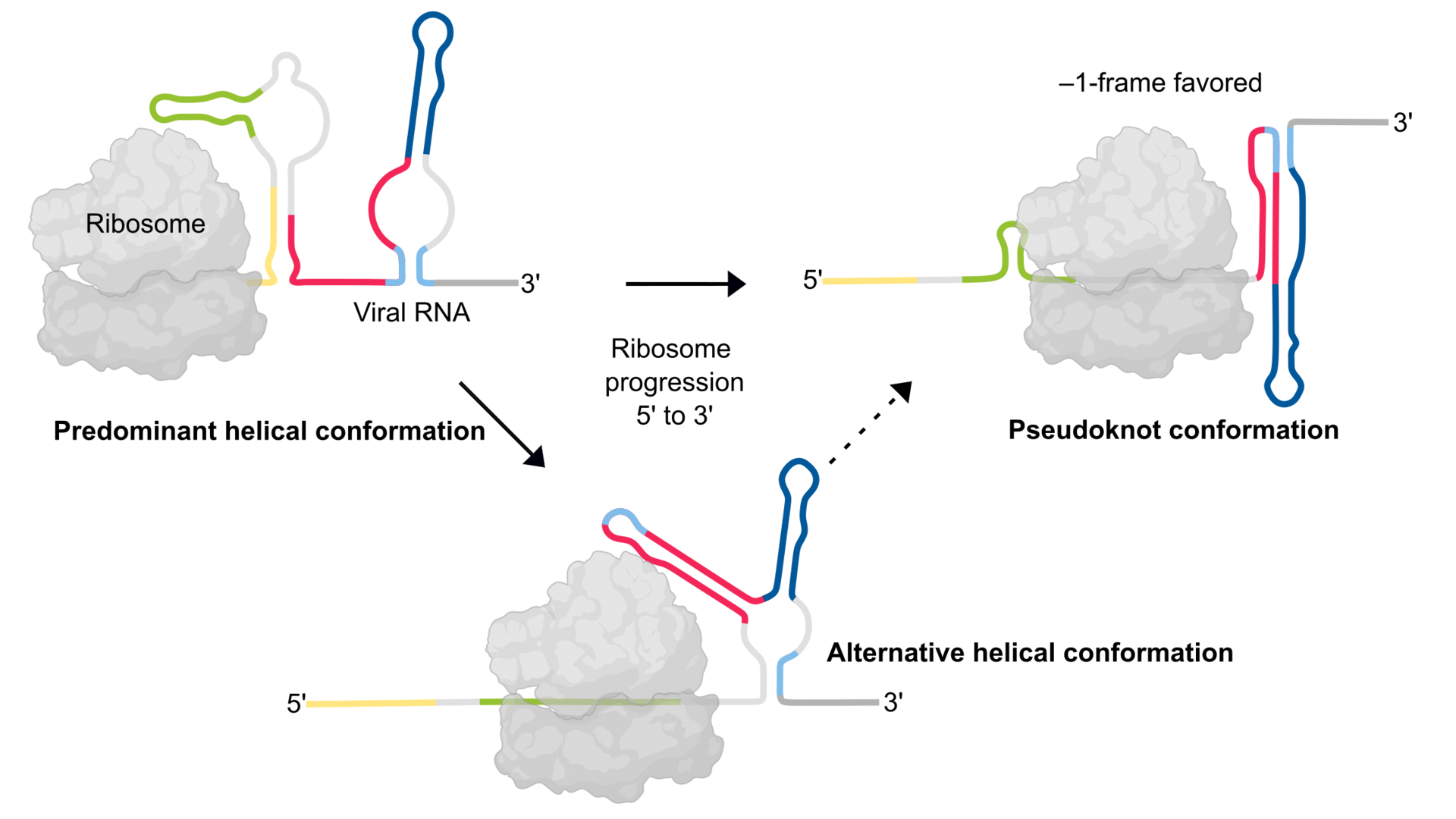
The current study revealed that a dynamic interplay of RNA structures governs the efficiency of frameshifting. The findings suggest that the pseudoknot structure of the SARS-CoV-2 frameshift RNA is essential for efficient frameshifting. However, it turned out that it is probably formed only transiently during translation. The alternative predominant structure consists of several helices, which do not allow the formation of the pseudoknot. As the ribosome translates the viral RNA, it has to sequentially unfold the upcoming RNA structures. When it starts translating the predominant helical form, at a certain moment, the part of the RNA region becomes available for pseudoknot formation, which then quickly forms.
“Multiple RNA structures work together to fine-tune SARS-CoV-2 gene expression,” explains Lukáš Pekárek, PhD student in the lab of Caliskan and first author of the study; “altering how the frameshifting RNA pairs with neighboring regions of the RNA can affect formation of various structures and, in turn, the ability of viruses to multiply in our cells.”
What's next?
Frameshift stimulatory elements in viral RNAs are evolutionary highly conserved. This makes them an attractive target for the development of new therapeutics, which would be less prone to the rise of resistance. Understanding the structure of the target RNA is thus essential. "So far, most of the attempts to discover novel therapeutics targeting the SARS-CoV-2 frameshift stimulatory element focused on the pseudoknot structure. While pseudoknot is the vital structure necessary for efficient frameshifting, our current study shows that it might not be an optimal target as the pseudoknot is formed only transiently during translation," says Neva Caliskan. Instead, shifting the focus towards targeting alternative stem-loop structures forming at the frameshift site can lead to the discovery of better hits, the researchers conclude.
Funding
The study received funding from the Helmholtz Association and grants from the European Research Council (ERC) [948636 to N.C.]; Helmholtz Young Investigator [VH-NG-1347 to R.S.].
Original publication
Pekarek L, Zimmer MM, Gribling-Burrer AS, Buck S, Smyth RP, Caliskan N (2023)
Cis-mediated interactions of the SARS-CoV-2 frameshift RNA alter its conformations and affect function
Nucleic Acids Research, DOI 10.1093/nar/gkac1184

Press contact