New review article
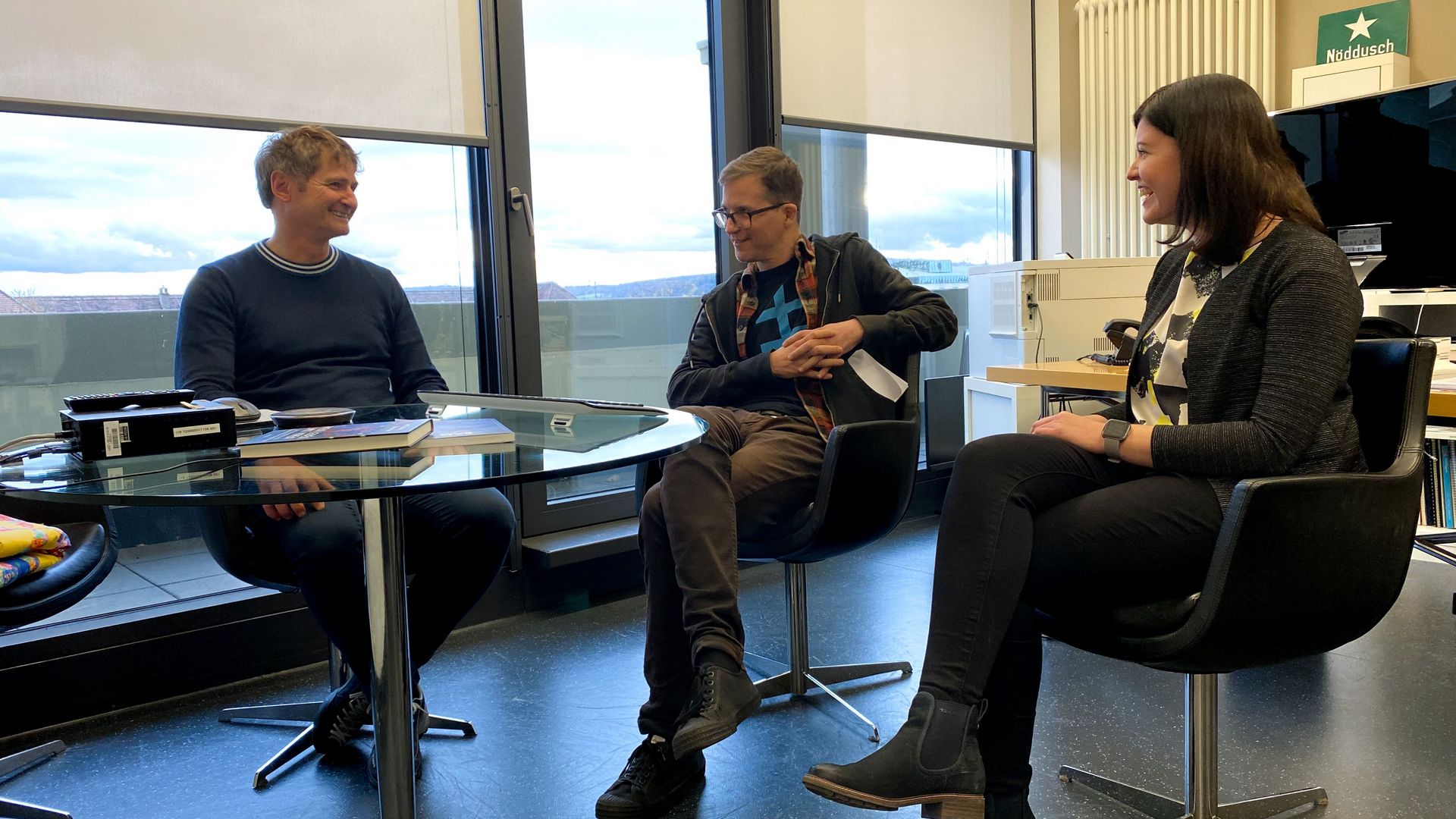
HIRI scientists discuss recent developments in transcriptome analysis of bacteria
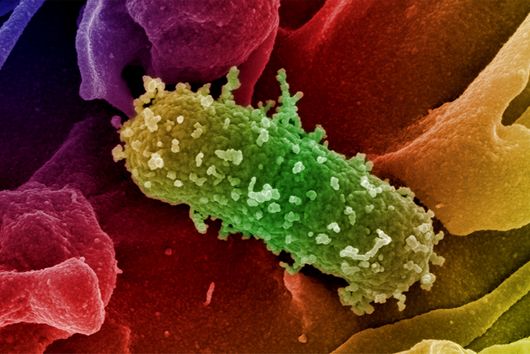
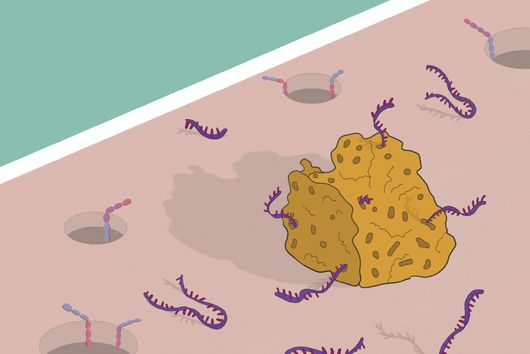
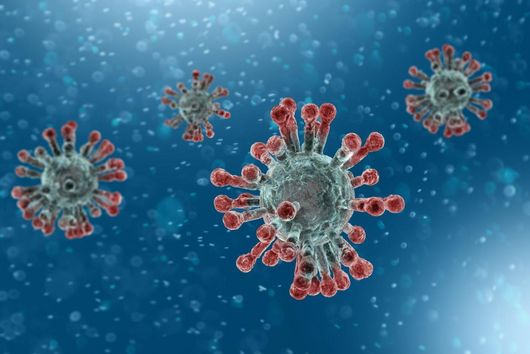
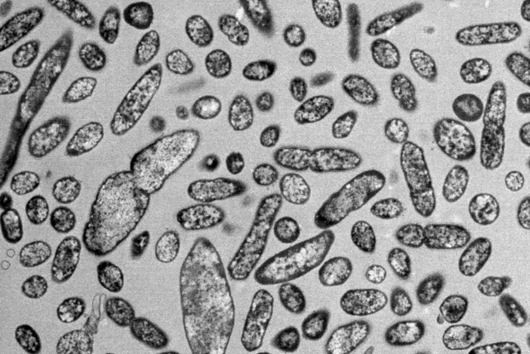
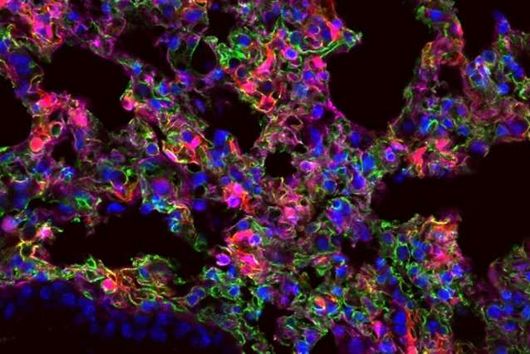
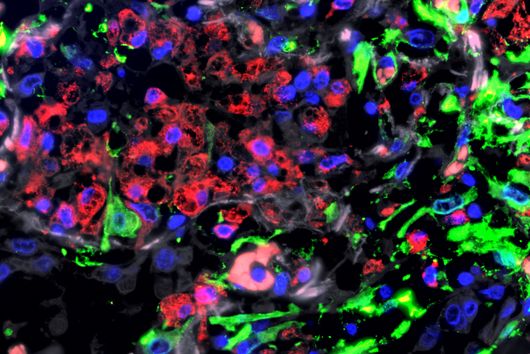
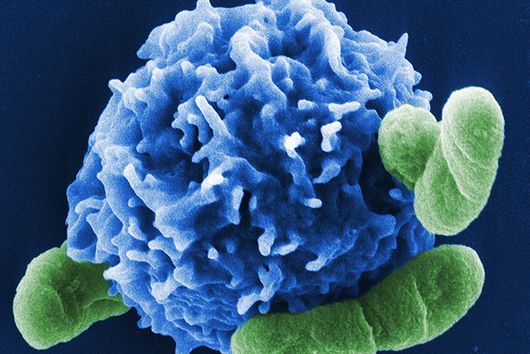
The sum of RNA molecules in a cell or tissue at a given state of development maps those gene sequences that are currently "switched on". This RNA profile of a cell is also known as the transcriptome and is the subject of research: With the aim of developing new therapeutic approaches, scientists want to understand at a molecular level how infections proceed.
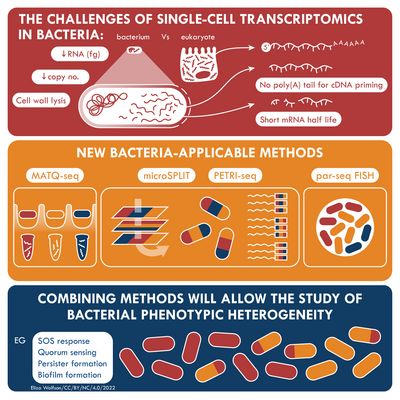
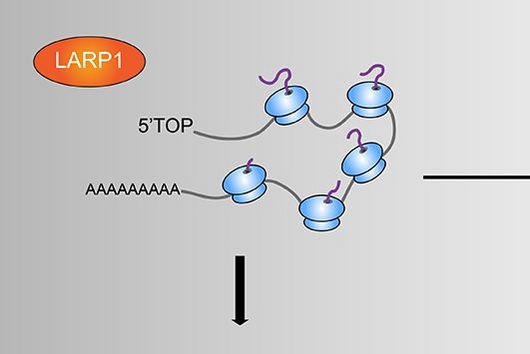
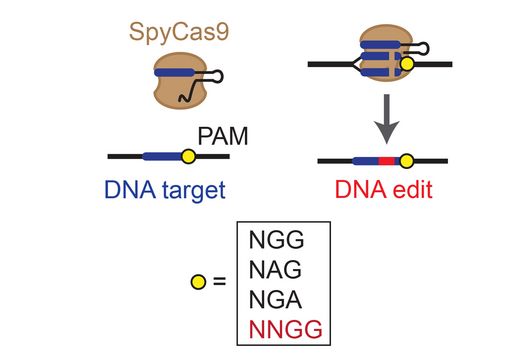
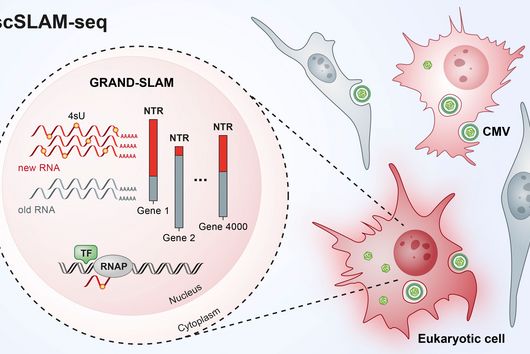
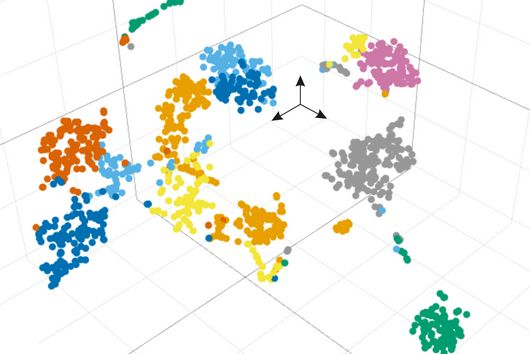
Transcriptome analysis of individual cells by single-cell RNA-seq (scRNA-seq) has become routine for eukaryotic tissues (cells of animals, plants and fungi). In contrast, developing methods to read the transcriptome of single bacterial cells has proven more challenging. Bacterial cells are harder to lyse, their RNA content is about two orders of magnitude lower than that of eukaryotic cells, and bacterial mRNAs are less stable than their eukaryotic counterparts. Most importantly, bacterial transcripts lack functional poly(A) tails, precluding simple adaptation of popular standard eukaryotic scRNA-seq protocols that come with the double advantage of specific mRNA amplification and concomitant depletion of rRNA.
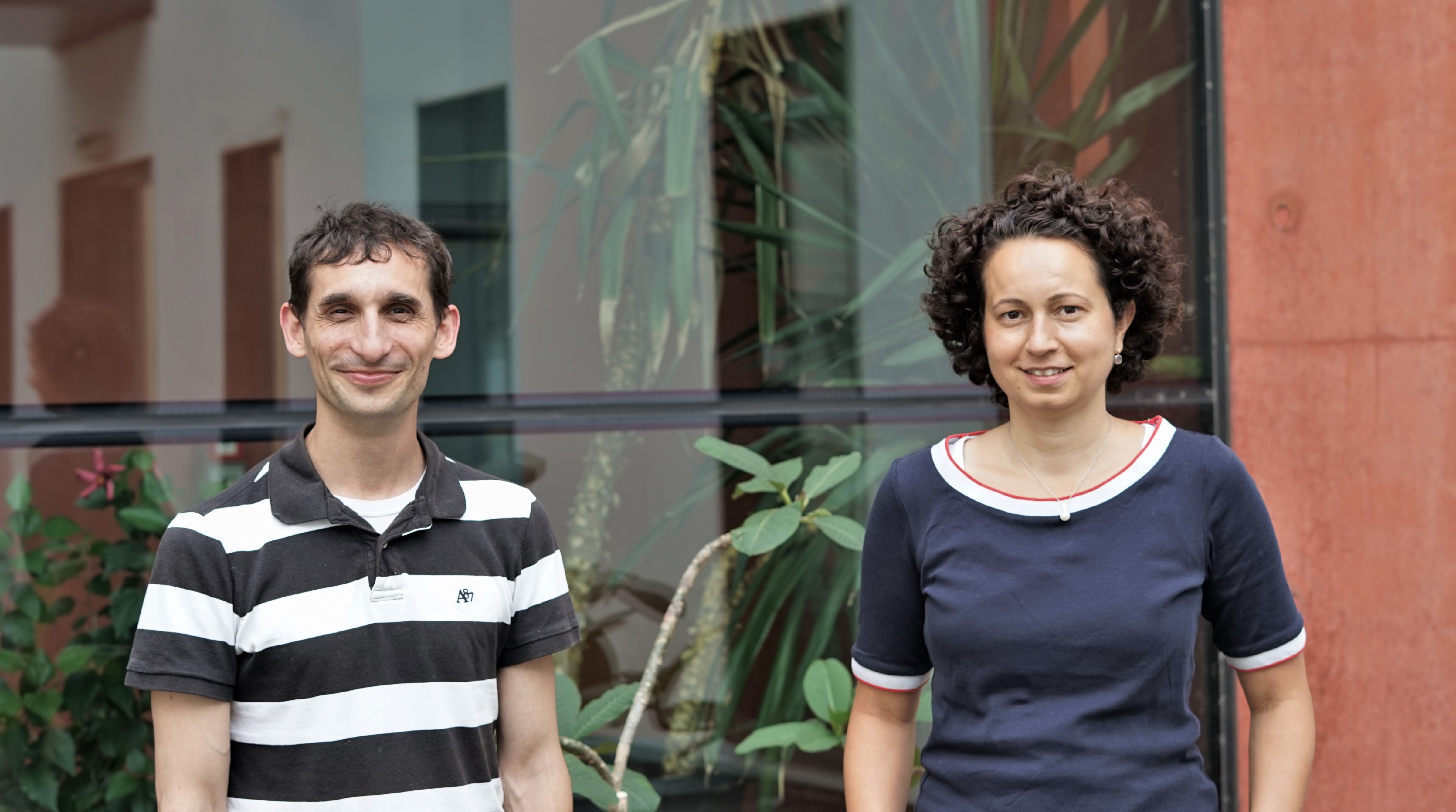
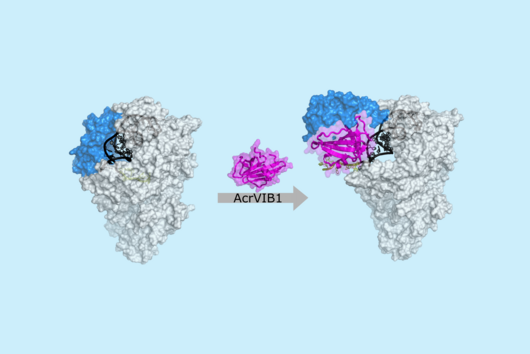
However, thanks to very recent breakthroughs in methodology, bacterial scRNA-seq is now feasible. In a review article that has recently been published in the journal microLife, Christina Homberger, Lars Barquist, and Jörg Vogel discuss several new approaches. They highlight the progress made and explain the technical challenges that still need to be overcome.